Chapter 28 Paramyxoviruses in Bats
Bats are an extraordinary group of mammals; not only are they the only mammals capable of sustained flight, but they are also found in almost all habitats, having a worldwide distribution, except for the highest mountains and extreme polar regions. Bats also occupy a diverse array of ecologic niches and contribute significantly to mammalian diversity, with more than 1000 species.39
DESCRIPTION OF PARAMYXOVIRUSES
There are two subfamilies and six genera within the family Paramyxoviridae. The genera Morbillivirus, Respiovirus, Rubulavirus, and Henipavirus make up the subfamily Paramyxovirinae, and the genera Pneumovirus and Metapneumovirus constitute the subfamily Pneumovirinae.56 Each of the six genera contains highly contagious human and animal pathogens.62
Paramyxoviruses have been found predominantly in mammals and birds, and most have a narrow host range in nature, but display a broad host cell range in culture.26 Transmission is generally horizontal, mainly through airborne routes, and no vectors are known.26 Primary replication is usually in the respiratory tract. Infection is generally cytolytic, but persistent infections often occur.56
Paramyxoviruses are pleomorphic and 150 to 300 nm in diameter.18 Virions are made up of a lipoprotein envelope and a nucleocapsid that surrounds a single strand of linear, negative-sense ribonucleic acid (RNA).13 Virion proteins common to all genera include three nucleocapsid-associated proteins: a nucleocapsid protein (N or NP), a phosphoprotein (P), and a large putative polymerase protein (L); and three membrane-associated proteins: an unglycosylated envelope protein (M) and two glycosylated envelope proteins, comprising a fusion protein (F) and an attachment protein (G or H or HN).56 The attachment and fusion proteins are of primary importance in inducing virus-neutralizing antibodies and immunity against reinfection.56,57 Antibodies to other viral proteins are also produced and some, nucleocapsid proteins in particular, are known to play a role as antigens for cytotoxic T cells.37
BATS AS VIRAL HOSTS
Historically, a wide range of viral infections, including flaviviruses, alphaviruses, rhabdoviruses, arenaviruses, reoviruses, and paramyxoviruses, have been identified in bats.60 More recently, a number of emerging zoonotic viruses have been detected in bats.32 These include Hantaan virus, isolated from the common serotine bat (Eptesicus serotinus) and the horseshoe bat (Rhinolophus ferrumepuinum) in Korea; Rift Valley fever virus, isolated from the bats Micropteropus pusillus and Hipposideros albae in the Republic of Guinea; a strain of yellow fever isolated from an Epomophorus Old World fruit bat in Ethiopia; and serologic evidence of Venezuelan equine encephalitis, St. Louis encephalitis, and eastern equine encephalitis viruses in bats in Guatemala.32 Although bat-variant rabies has long been recognized in the United States, the prevalence of human rabies cases attributed to that variant has increased in recent years.46 Most recently, strong evidence shows that horseshoe bats (Rhinolophus spp.) may be the source of the severe acute respiratory syndrome (SARS) coronavirus.40,42 Also, Old World fruit bats of the genera Hypsignathus, Epomops, and Myonycteris may be natural hosts of Ebola virus, as found in Gabon and Republic of Congo.41
Bats may travel hundreds of kilometers (or miles) in a matter of days. Besides having significant implications for disease spread, this also suggests that populations of pathogens carried by bats are likely to be relatively homogenous across wide geographic areas.9 Studies of Old World fruit bats using satellite telemetry have shown that individuals can travel more than 2000 km in 1 year and traverse significant bodies of open sea, such as the Torres Strait between Australia and New Guinea and the Strait of Malacca between peninsular Malaysia and Sumatra (www.henipavirus.com)10,61 Figure 28-1. These long-distance movements may transmit pathogens over great distances and enable exchange between bat populations on different land masses.
Population size and density are positively associated with the diversity of pathogens hosted by mammalian species.2 A large population size, as seen for many colonial bats, supports pathogen reproduction by providing a constant supply of individuals susceptible to infection and thus allows persistence of the pathogen.3
Regular, but not constant, contact between individual bats from different subpopulations allows for partial connectivity between colonies of bats. A metapopulation may exist where a spatial mosaic involves a constellation of subpopulations of which, at any given time, some are susceptible, some infected, and some immune to a particular disease.43 This is beneficial for genetic diversity and may permit pathogens, particularly viruses, to persist in a species with a total population that would otherwise be too small to maintain the disease.8 This results in these species having considerable potential to act as vectors for, and disseminators of, viruses and other pathogens.
Some authors have proposed that bats are unique in their response to viral infection and are able to sustain viral infections without disease.60 However, many other small mammals act as reservoirs for viruses without evidence of disease, and recent analysis of a database on all emerging infectious diseases of humans suggests that bats (which represent as much as a quarter of all mammalian species) do not harbor a disproportionate number of the known emerging zoonotic viruses.71
PARAMYXOVIRUSES OF CHIROPTERA
Hendra Virus
In September 1994, an outbreak of severe respiratory disease of horses occurred in the Brisbane suburb of Hendra in eastern Australia (Queensland)48 Figure 28-2. The index case was a pregnant Thoroughbred mare, and 16 other horses at two sites showed signs of loss of appetite, dyspnea, and copious frothy nasal discharge. Twelve of the affected horses died a few days after the onset of signs.7 Two people who had close contact with the index case also became infected. One of them, a stable worker, developed flulike signs and recovered. The other person, a horse trainer, showed rapid development of respiratory illness and died 11 days later.58
It was thought that the pattern of disease in the horses reflected a point source of infection, and that all other cases were a direct result of transmission from the pregnant mare.7 A range of pathogens and toxins were investigated and excluded from the diagnosis. A novel virus was cultured from the lungs of five of the affected horses and from the kidneys of the fatal human case.49 The virus showed characteristics suggesting it belonged to the family Paramyxoviridae, although there was minimal cross-reactivity between the virus and a range of antisera to other paramyxoviruses. The virus showed 50% homology of the partial M protein gene sequence of several morbilliviruses and thus was initially called equine morbillivirus (EMV).48 The name was subsequently changed to Hendra virus (HeV) when it became apparent that horses were not the natural host for the virus and that it did not belong in the genus Morbillivirus.
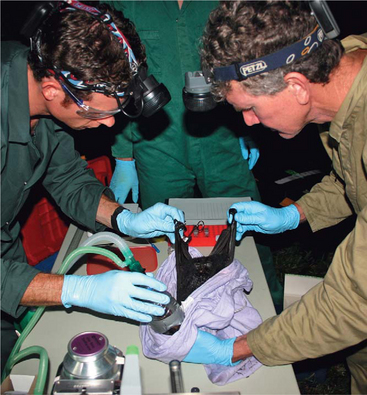
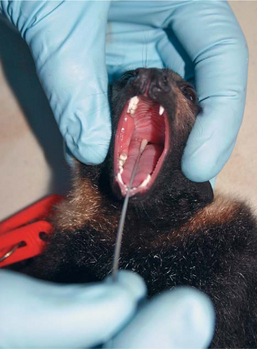
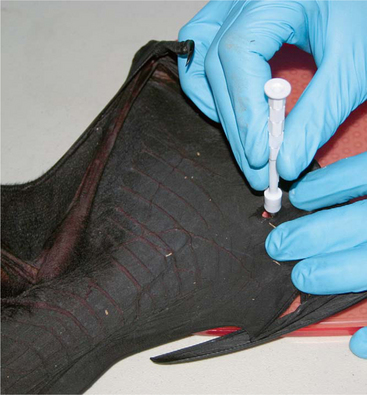
Surveillance of wildlife species identified flying foxes (genus Pteropus, family Pteropodidae) as the likely natural host of the virus. The infection was found to be widespread in four of the flying fox species found on mainland Australia: the black flying fox (Pteropus alecto), gray-headed flying fox (P. poliocephalus), little red flying fox (P. scapulatus), and spectacled flying fox (P. conspicillatus).22,29 Sampling of 46 species of ground-dwelling mammals revealed no evidence of HeV exposure.69 Studies of seroprevalence of HeV antibodies in flying foxes in Australia indicate a prevalence of approximately 50%25 Figures 28-3 through 28-5.
Since the first outbreak, further outbreaks of HeV have occurred in Queensland, including Mackay in 1994, involving fatal infections of both horses and a human; Cairns in 1999, involving a single horse; and Cairns and Townsville in 2004, involving both horses and a veterinarian.23
Hendra virus infection of terrestrial mammals, including humans, results in a systemic vasculitis with significant pathology of the lung and central nervous system (CNS).34,66,70 Viral antigen is detected in vascular endothelium and frequently recovered from nasopharangeal swabs, urine, and internal organs, including lung and brain.19,34 Experimental HeV infection of flying foxes, however, appears to cause only a sporadic subclinical vasculitis, even at infective doses lethal to horses.68,69 Viral antigen is detected in the tunica media rather than endothelial cells, which may help explain why flying foxes appear to be spared from clinical disease.20 Experimental infection of flying foxes has also shown placental transfer of the virus to a fetus.34
Nipah Virus
Nipah virus (NiV) was first described in March 1999 in the investigation of an outbreak of disease in pigs and humans in Malaysia (see Figure 28-2). In the course of the outbreak, 265 humans were infected, 105 fatally.15 Infected pigs were identified as the primary source of human infection, and over 1 million pigs were culled to control the outbreak. Wildlife surveillance identified the Malayan flying fox (Pteropus vampyrus) and island flying fox (Pteropus hypomelanus) as probable natural hosts of NiV.38
Subsequent studies have also found serologic evidence of NiV infection in the Malayan flying fox, island flying fox, and Lyle’s flying fox (Pteropus lylei) in Thailand, in Lyle’s flying fox in Cambodia, and in the Indian flying fox (Pteropus giganteus) in Bangladesh.35,51,63 NiV has strong serologic and sequence similarities to HeV and is the second member of the genus Henipavirus.64
Subsequent to NiV’s emergence in Malaysia, five outbreaks of NiV-associated disease in humans were described in Bangladesh between April 2001 and February 2005.4–635 As of 11 February 2005, a total of 122 cases had been recognized by the Bangladesh Directorate of Health Services, at least 78 (64%) of which were fatal. A number of the characteristics of the Bangladesh outbreaks were similar to the outbreak in Malaysia: delayed recognition, a primary presentation of humans with fever and CNS signs, and a high case-fatality rate. In marked contrast to the Malaysian outbreak, however, infection in humans was not associated with disease in pigs, and evidence indicated horizontal human transmission. Further, the pattern of the Bangladesh outbreaks suggests a sporadic, geographically scattered introduction of infection to humans. Nucleotide sequence data also support a different epidemiology in Bangladesh. Data obtained from human cases in Malaysia suggest a single source of human infection from the porcine amplifying host.1,11,15 Data from Bangladesh cases formed a cluster clearly distinct from the Malaysian sequences, but differed from each other by approximately 0.8%, suggesting possibly multiple introductions of virus into humans.30
The pathologic effects of NiV in terrestrial mammals are similar to those of HeV, with infection resulting in a systemic vasculitis and significant pathology of the lung and CNS.34,66 In contrast to HeV, however, viral antigen is often found in bronchial and alveolar epithelium. NiV has not been associated with clinical disease in flying foxes.20
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree