Picornaviridae
Abstract
This chapter describes the properties of picornaviruses and features of the diseases they cause in animals.
Keywords
Picornavirus; Aphthovirus; Cardiovirus; Enterovirus
Picornaviruses such as foot-and-mouth disease virus and poliovirus have played an important role in the histories of both human and veterinary medicine and virology. In 1897, Loeffler and Frosch showed that foot-and-mouth disease was caused by an agent that passed through filters which held back bacteria; this was the first demonstration that a disease of animals was caused by a filterable virus. Poliovirus, the cause of human poliomyelitis, was not identified until some 10 years later. Poliovirus and other picornaviruses were also involved in key developments in the techniques used to study viruses, including the growth of viruses in cell culture, quantitative plaque assays, infectious clones of specific viruses, X-ray crystallographic analysis of virion structure at the atomic level, RNA replication, and viral protein synthesis. The development of poliovirus vaccines in the 20th century has greatly reduced the occurrence of human poliomyelitis, a once prevalent and often devastating disease that has been recognized since antiquity. Indeed, the advent of highly effective inactivated poliovirus vaccines has stimulated efforts to eradicate the disease from the global human population, as was done for smallpox.
In the second half of the 19th century and the first half of the 20th century, repeated rapidly spreading epidemics of foot-and-mouth disease resulted in great losses, as increasingly intensive systems of livestock production were developed in many countries. Producers demanded of their governments control programs to deal with these epidemics, as well as programs to prevent reintroductions. For example, in 1884, the United States Congress created the Bureau of Animal Industry within the Department of Agriculture. Its principal mission was to deal with foot-and-mouth disease and two other diseases, contagious bovine pleuropneumonia and hog cholera (classical swine fever). From its beginning, this agency pioneered the development of veterinarians with special skills in disease control. An extensive epidemic of foot-and-mouth disease in 1914 accelerated the creation of disease control programs and the training of more specialized veterinarians. Eventually, this evolved into the complex field- and laboratory-based systems needed to assure the freedom of domestic livestock industries from foreign animal diseases. Similar developments occurred in other countries with increasingly intensive livestock industries, in each case advancing the scope of the veterinary medical profession from its roots in equine medicine and surgery.
Properties of PICORNAVIRUSES
Classification
The family Picornaviridae is now included with the families Dicistroviridae, Iflaviridae, Marnaviridae, and Secoviridae in the Order Picornavirales. All viruses within this order share the following features: (1) conserved RNA-dependent RNA polymerase; (2) a genome that has a protein (VPg) attached to the 5′ end; (3) absence of overlapping open reading frames within the genome; and (4) viral RNA translated into a polyprotein before processing. Viruses within the families Iflaviridae, Marnaviridae, and Secoviridae infect only insects, plants, or algae and will not be considered further in this chapter. Viruses in the family Dicistroviridae infect insects and crustaceans, and the disease, designated as “Taura syndrome,” has resulted in devastating losses in shrimp farms.
The family Picornaviridae has undergone a significant expansion in recent years, due principally to the identification of previously unknown picornaviruses by the “next-generation” sequencing of clinical and environmental samples. The family is divided currently into 29 genera, of which 23 include only a single virus species. In addition to the well-established genera of Aphthovirus, Enterovirus, Teschovirus, Cardiovirus, Erbovirus, Kobuvirus, Hepatovirus, and Parechovirus, the identification and comparative analysis of new and existing picornavirus genomes has resulted in the creation of 21 new genera: Aquamavirus, Avihepatovirus, Avisivirus, Cosavirus, Dicipivirus, Gallivirus, Hunnivirus, Kunsagivirus, Megrivirus, Mischivirus, Mosavirus, Oscivirus, Pasivirus, Passerivirus, Rosavirus, Sakobuvirus, Salivirus, Sapelovirus, Senecavirus, Sicinivirus, and Tremovirus. The genus Enterovirus is the largest genus within the family and contains viruses with most relevance to human medicine; included in this genus are enteroviruses that use the gastrointestinal tract as the primary site of replication (eg, polio-, echo-, and coxsackie viruses), as well as the rhinoviruses that infect the upper respiratory tract. With the exception of the aphthoviruses that are yet to be changed, picornavirus species have been renamed recently to remove host species names that have been replaced with alphabetical assignments. Given the seemingly ever-changing and potentially confusing taxonomic organization of picornaviruses, coupled with the fact that taxonomic assignments do not consistently correlate with the biological behavior of individual picornaviruses (including the nature of the disease they induce in animals, if any), this chapter will be organized according to animal species rather than the taxonomic assignment of each virus. Important picornaviruses in human and veterinary medicine are listed in Table 26.1.
Table 26.1
Genera and Species of Picornaviruses Causing Important Diseases of Animals and Humans
| Genus | Virus Species | Hosts Affected | Disease/Comments |
| Aphthovirus | Foot-and-mouth disease viruses | Cattle, sheep, swine, goats, wildlife ruminant species | Foot-and-mouth disease; 7 serotypes |
| Equine rhinitis A virus | Horses, camelids | Systemic infection with respiratory signs | |
| Bovine rhinitis A virus | Cattle | Mild respiratory signs | |
| Bovine rhinitis B virus | Cattle | Mild respiratory signs | |
| Avihepatovirus | Avihepatovirus A [Duck hepatitis A virus]a | Duck | Hepatitis |
| Cardiovirus | Cardiovirus A [Encephalomyocarditis virus] | Rodents, swine, elephants, primates, mammals in contact with rodents | Encephalomyelitis and myocarditis in swine and elephants; rarely in other species; 2 serotypes |
| Cardiovirus B [Theilovirus] | Rodents | Murine poliomyelitis; 15 genotypes | |
| Cardiovirus C [Boone cardiovirus] | Rats | Fecal isolation | |
| Enterovirus | Enterovirus A [Human enterovirus A] | Human, simian | Hand, foot-and-mouth disease, meningitis; 25 serotypes |
| Enterovirus B [Human enterovirus B] | Humans | Rash, respiratory disease, paralysis; 61 serotypes | |
| [Swine vesicular disease virus] | Swine | Vesicular disease | |
| Enterovirus C [Human enterovirus C] | Human | Poliomyelitis, Respiratory disease; 23 serotypes | |
| Enterovirus D | Humans, primates | Respiratory disease, focal limb paralysis; 5 serotypes | |
| Enterovirus E [Bovine enterovirus group A] | Cattle | Asymptomatic or mild enteric, respiratory, reproductive disease; 4 types | |
| Enterovirus F [Bovine enterovirus group B] | Cattle | Asymptomatic or mild enteric, respiratory, reproductive disease; 6 types | |
| Enterovirus G [Porcine enterovirus B] | Swine, ovine | Usually asymptomatic infection; 11 serotypes | |
| Enterovirus H | Simian | Usually asymptomatic infection | |
| [Simian enterovirus A] | |||
| Enterovirus J | Simian | Usually asymptomatic infection; 6 types | |
| Rhinovirus A, B, and C [Human rhinovirus A, B, and C] | Humans | Respiratory disease; 80 (A), 32 (B), and 54 (C) serotypes | |
| Erbovirus | Erbovirus A [Equine rhinitis B virus] | Equine | Associated with mild rhinitis |
| Kobuvirus | Aichivirus A [Aichi virus] | Humans, canine, feline, rodents, birds | Gastroenteritis, enteric infections |
| Aichivirus B [Bovine kobuvirus] | Ovine, bovine, ferret | Detected in feces and serum | |
| Aichivirus C [Porcine kobuvirus] | Swine | Fecal detection | |
| Megrivirus | Melegrivirus A | Chickens, turkeys, ducks | Fecal detection, associated with hepatitis in turkeys |
| Sapelovirus | Avian sapelovirus | Ducks | Hepatitis |
| Sapelovirus A [Porcine sapelovirus/porcine enterovirus A] | Swine | Diarrhea | |
| Sapelovirus B [Simian sapelovirus] | Simian | ||
| Teschovirus | Teschovirus A [Porcine teschovirus] | Swine | Type 1; encephalomyelitis |
| Types 2–13; mild/asymptomatic | |||
| Tremovirus | Tremovirus A [Avian encephalomyelitis virus] | Chicken, pheasant, turkey, quail | Encephalomyelitis |
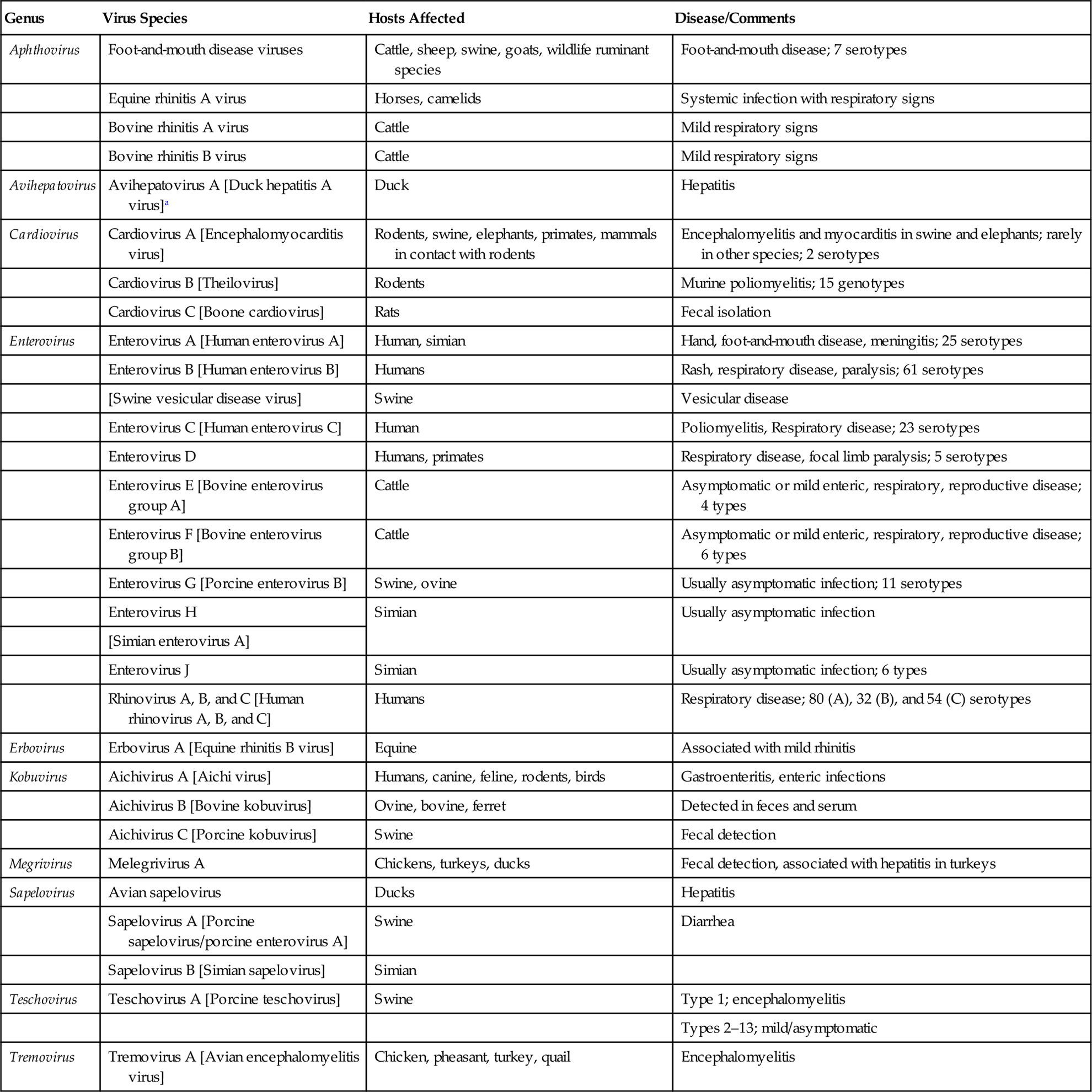
aParentheses[] indicate former (and often commonly used) names of individual viruses.
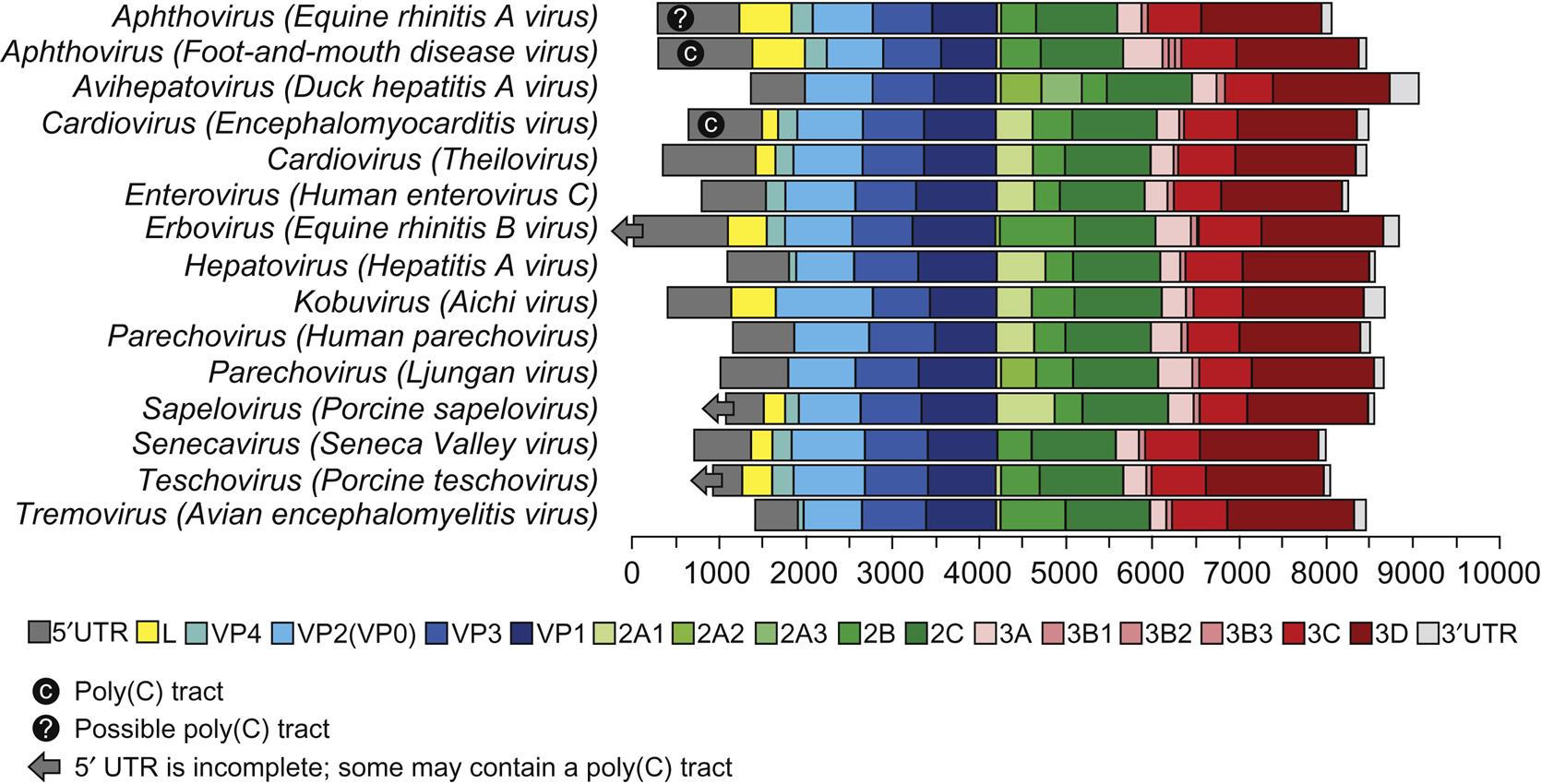
A significant difference among viruses in the various picornavirus genera is their stability at low pH; such differences were utilized in the classification of picornaviruses before molecular techniques were available. Specifically, the aphthoviruses are unstable below pH 7, whereas the enteroviruses, hepatoviruses, cardioviruses, and parechoviruses are stable at pH 3. However, other major similarities and differences were identified with the availability of complete genomic sequence data. All picornaviruses are single-stranded, positive-sense RNA viruses with a 5′-untranslated region (5′-UTR). The RNA is uncapped, but with a viral protein (VPg) covalently linked to the 5′ end. There are major structural differences in the 5′-UTR among the genera of the picornavirus family: the length of the 5′-UTR in picornaviruses varies from approximately 500 to 1200 nt and contains one of at least five different internal ribosome entry sites (IRESs). Cardioviruses, aphthoviruses, erboviruses, kobuviruses, teschoviruses, and sapeloviruses are among the 16 genera that are also distinguished by the presence of a leader protein (L) encoded upstream of the capsid proteins (Fig. 26.1). Foot-and-mouth disease virus, aquamavirus A, mosavirus A, and possibly passerivirus A all have multiple (2–3), but not identical VPg proteins that are present in equimolar amounts among the virion RNAs. Other virus species within each genus do not necessarily contain multiple copies of VPg.
Virion Properties
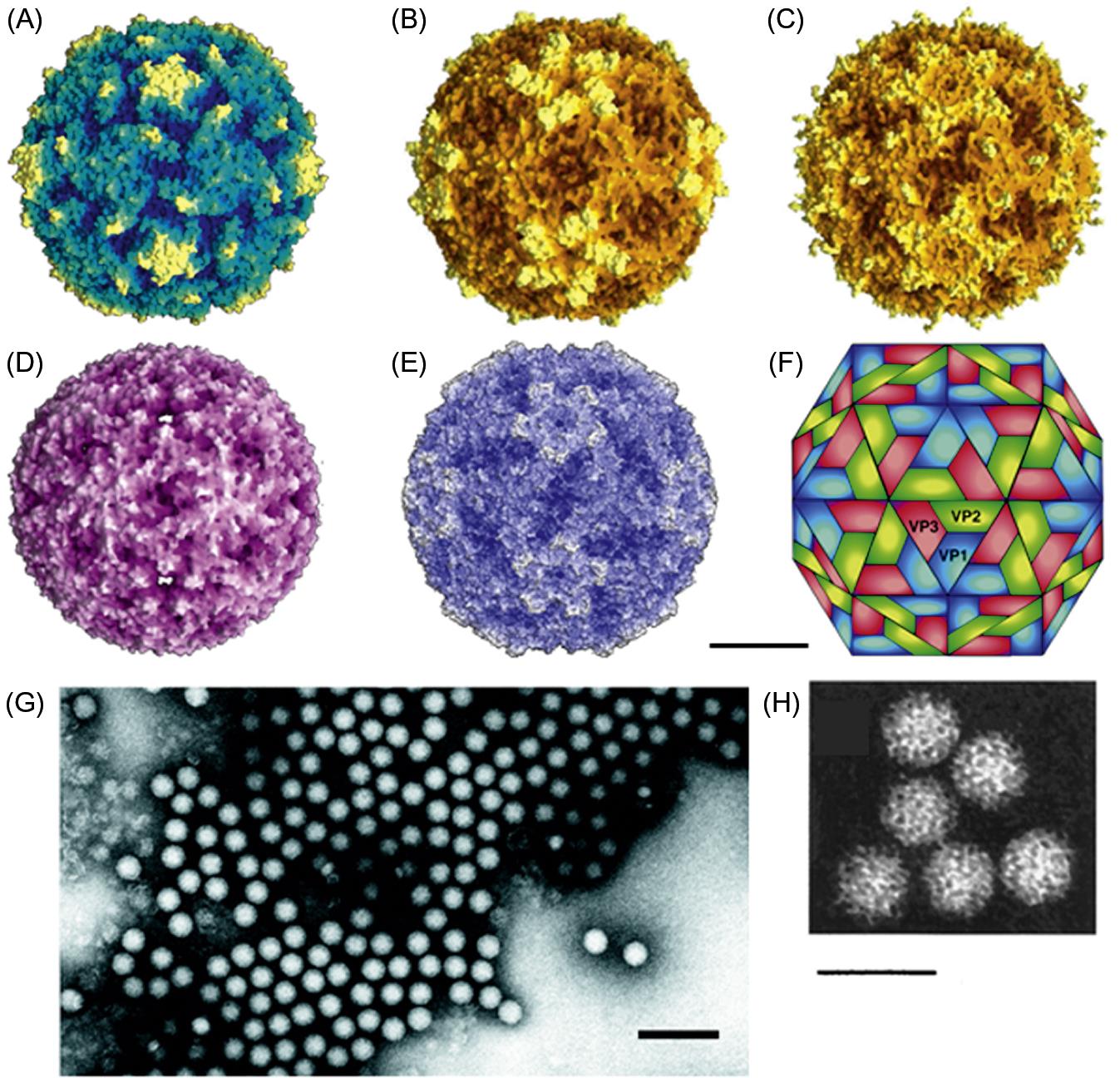
Picornavirus virions are nonenveloped, approximately 30 nm in diameter, and have icosahedral symmetry (Fig. 26.2; Table 26.2). Virions appear smooth and round in electron micrographs and in images reconstructed from X-ray crystallographic analyses. The genome consists of a single molecule of linear, positive-sense, single-stranded RNA, 7–8.8 kb in size. Both the 5′ and 3′ ends of the RNA contain untranslated regulatory sequences. The genomic RNA is polyadenylated at its 3′ end and has a protein, VPg, linked covalently to its 5′ end. Genomic RNA is infectious.
Table 26.2
Virions appear smooth and round in outline, are nonenveloped, <30 nm in diameter, and have icosahedral symmetry
The genome consists of a single molecule of linear, positive-sense, single-stranded RNA, 7–8.8 kb in size
Genomic RNA is polyadenylated at its 3′ end and has a protein, VPg, linked covalently to its 5′ end; genomic RNA is infectious
Virion RNA acts as mRNA and is translated into a polyprotein(s), which is then cleaved to yield some 11 or 12 individual proteins
Cytoplasmic replication
Picornavirus virions are constructed from 60 copies each of four capsid proteins, VP1 (also designated 1D), VP2 (1B), VP3 (1C) (Mr approximately 30,000 for each), and VP4 (1A) (Mr 7000–8000), and a single copy of the genome-linked protein, VPg (3B) (Mr variable). The exceptions to the “four capsid protein” rule are parechoviruses and kobuviruses, in which the VP0, a polyprotein that includes VPs 2–4, remains uncleaved. VP1, VP2, and VP3 are structurally similar to one another, each being composed of a wedge-shaped, eight-stranded β-barrel and differing primarily in the size and conformation of the loops that occur between the strands and also in the extensions of their amino and carboxyl termini. Amino acid substitutions correlating with antigenic variation occur in the surface-oriented loop regions. For foot-and-mouth disease virus, at least five antigenic sites have been identified. The VP1 proteins are located around the fivefold axes of icosahedral symmetry, and VP2 and VP3 alternate around the two- and threefold axes. The amino terminal extensions of these three proteins form an intricate network on the inner surface of the protein shell. The small, myristoylated protein, VP4, is located entirely at the inner surface of the capsid, probably in contact with the RNA.
In poliovirus and rhinovirus virions, the packing together of VP1, VP2, and VP3 results in the formation of a “canyon” around the fivefold axes of the virion. The amino acids within the canyon, particularly those on the canyon floor, are conserved, but the amino acids on the “rim” of the canyon are variable. For polio- and rhinoviruses, the conserved amino acids on the floor of the canyon form the points of attachment of the viruses to cell surface receptors. Changes on the rim of the canyon affect the binding affinity to the receptor. Beneath the floor of the canyon in picornaviruses is a hydrophobic pocket accessible from the surface via a small opening. This pocket is a target for chemotherapeutic drugs that may block capsid changes critical for effective receptor binding or RNA release. Foot-and-mouth disease viruses have a comparatively smooth surface, with no canyon structure. The attachment site for host-cell receptors is located on VP1, within the G–H loop. These sites have serotype and subtype antigenic specificities that differ among the various strains of foot-and-mouth disease virus (see Fig. 1.5 for more information on capsid protein structure).
The stability of picornaviruses to environmental conditions is important to the epidemiology of the diseases they cause, and in the selection of methods of disinfection. For example, if protected by mucus or feces and shielded from strong sunlight, most picornaviruses are relatively heat stable at usual ambient temperatures. Some enteroviruses, for instance, may survive for several days, and often weeks, in feces. Aerosols of foot-and-mouth disease virus are less stable, but under conditions of high humidity they may remain viable for several hours. Because of differences in their pH stability, only certain disinfectants are suitable for use against each virus; for example, sodium carbonate (washing soda) is effective against foot-and-mouth disease virus, but is not effective against swine vesicular disease virus.
Virus Replication
The replication of picornaviruses is described in detail in Chapter 2: Virus Replication, see Fig. 2.8. Poliovirus (genus Enterovirus), which in nature only infects humans and nonhuman primates, has historically been the principal model for studying the replication of RNA viruses. This model served as the basis for analyzing the replication pattern for all other picornaviruses, and deviations from this model have provided support for the continuing reclassification of the picornaviruses.
The cellular receptors for many picornaviruses are known, and are surprisingly diverse (Table 2.1). The receptors for polioviruses, coxsackie B viruses, and some human rhinoviruses are members of the immunoglobulin (Ig) superfamily. For other picornaviruses, many other cell surface molecules serve as receptors and coreceptors, including heparan sulfate, low-density lipoproteins, extracellular matrix-binding proteins, and integrins. Foot-and-mouth disease virus can use two different receptors, depending on the passage history of the individual virus strain; specifically, field strains of foot-and-mouth disease virus bind to integrins, whereas cell-culture-passaged virus can use heparan sulfate as a receptor, but this change in receptor specificity results in attenuation of the virus. Foot-and-mouth disease viruses can also enter cells via Fc receptors if virions are complexed with nonneutralizing IgG molecules. This pathway, termed the antibody-dependent enhancement of infection pathway, is of unknown significance, but may be important in the long-term carrier state that occurs in certain ruminants.
The pathway(s) following attachment of the virus to its receptor and the release of the virion RNA into the cytoplasm of the host cell varies among the picornaviruses. The specific pathways used may reflect the pH stability of the virions of picornaviruses in different genera. For poliovirus, its interaction with the cell receptor induces structural changes in the virion such that VP4 is released from the virion and the amino terminal of VP1 shifts from the interior of the virion to the surface. This amino terminal region is hydrophobic and can induce membrane permeability by the generation of a pore in the cell membrane. Both pH and VP4 myristoylation influence the membrane permeability induced by the formation of VP4 associated pores. It is proposed that the RNA from the virus particle gains access to the cytoplasm of the host cells through this membrane pore (see Fig 2.5). The endosomal pathway of virus entry is not utilized by poliovirus, as indicated by the lack of inhibition of infectivity by drugs that block this pathway. For foot-and-mouth disease virus, the entry process is different, as the binding of this virus to its receptor does not induce changes in the virion structure, but simply functions as a docking mechanism; once bound to the receptor, the virus enters the cell through the clathrin-dependent endosomal pathway. Weak bases that increase the pH of the endosomes block replication of foot-and-mouth disease virus; low pH of the endosome induces the capsid to disassociate into pentamers, with release of the viral RNA. In contrast, poliovirus, which is stable at low pH, cannot utilize the low pH environment of the endosome for penetration and release of the virion RNA in the same manner as foot-and-mouth disease virus.
After adsorption, penetration, and intracellular uncoating, VPg at the 5′ end of the RNA is removed from the virion RNA by cellular enzymes (see also Fig. 2.8). Picornaviruses have evolved a cap-independent mechanism of translation that permits normal cellular cap-dependent translation to be inhibited by viral gene products. Initiation of translation does not proceed by the well-established Kozak scanning model. Instead, ribosomal binding to viral RNA occurs in a region of the 5′-UTR of the genome known as the internal ribosome entry segment (IRES). This segment of the viral genomic RNA is folded into cloverleaf-like structures, which bind specifically to host-cell proteins that play key roles in initiating the synthesis of viral protein and RNA. The IRES has been recognized as a determinant of the phenotype and neurovirulence of some viruses. At least five different IRES structures have been identified in the picornavirus family. Cadicivirus A (Canine dicistronic picornavirus), in the genus Dicipivirus, is the only picornavirus with a dicistronic genome, containing an IRES in the usual location in the 5′ UTR, as well as another in a unique intergenic region that occurs between the P1 (structural protein coding region) and P2 regions (described in the next paragraph). A similar virus recently was identified in mice. All picornavirus genomes have a 3′-UTR and a 3′-poly(A) tail that is encoded by the virus rather than added by host-cell polyadenylation enzymes. The function of the poly(A) tails is not known, but removal of poly(A) tails from the virion RNA renders it noninfectious. A short (~48 nucleotide) “bar bell”-like RNA structure has been identified in the 3′-UTR of sicinivirus, passerivirus, gallivirus, avihepatovirus and some kobuviruses. It has been suggested that the 3′-UTR is involved in the regulation of RNA synthesis, but is not essential for infectivity.
The RNA genome of picornaviruses (with the exception of cadicivirus A) comprises a single open reading frame that is translated into a single polyprotein (Fig. 26.2). The polyprotein is cleaved posttranslationally by virus-encoded proteinases in a stepwise fashion to produce 11 or 12 proteins. Cadicivirus A is unique in that it contains 2 functional IRES elements and 2 open reading frames, encoding 2 polyprotein precursors separated by an intergenic region of 588 bases. In all picornaviruses some of the intermediate cleavage products have functions vital to replication. The 5′-terminal region of the genome encodes the virion proteins VP0 (VP4+VP2), VP3, and VP1, in that order; the common designation of these proteins for foot-and-mouth disease virus is respectively 1A, 1B, 1C, and 1D. Viruses in 16 genera of the family have a nonstructural L protein sequence at the 5′ end of the coding sequence. Foot-and-mouth disease virus has two initiation codons at the 5′ end of the genome, which results in two forms of the L protein. The function of the alternative forms of the L protein is not known, but the two codons are strictly conserved in all isolates, and removal of the L coding region in foot-and-mouth disease virus produces an attenuated virus. The middle region of the genome of picornaviruses encodes nonstructural viral proteins (designated 2A, 2B, and 2C) that can exhibit protease activity in viruses within some genera. The 3′ end of the genome encodes additional nonstructural proteins 3A, 3B, 3C, and 3D. Protein 3C has invariably a protease activity, 3B is the virion protein VPg, and 3D is the core RNA polymerase.
Viral RNA synthesis takes place in a replication complex, which comprises RNA templates, the virus-coded RNA polymerase, and several other viral and cellular proteins, tightly associated with newly assembled smooth cytoplasmic membrane structures. Synthesis of the complementary strand is initiated at the 3′ terminus of the virion RNA and uses the uridylated protein, VPg, as a primer. The completed complementary strand in turn serves as a template for the synthesis of virion RNA. Most of the replicative intermediates found within the replication complex consist of a full-length complementary (negative-sense) RNA, from which several nascent plus-sense strands are transcribed simultaneously by viral RNA polymerase.
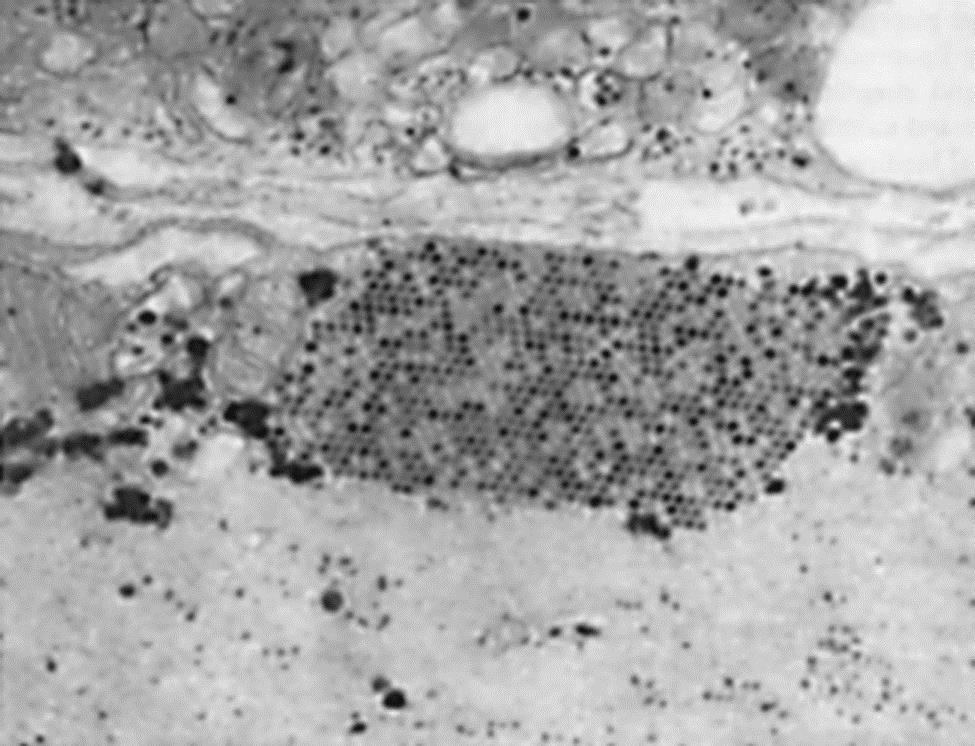
With the absence of a 5′ cap on picornavirus mRNAs, these viruses have evolved unique mechanisms for inhibiting the translation of cellular mRNAs. The 2A protease of poliovirus or the L protease of foot-and-mouth disease virus cleaves the eIF4G of the translation initiation complex in a manner that permits viral mRNA to be preferentially translated. Conversely, encephalomyocarditis virus (now designated as cardiovirus A) blocks phosphorylation of a protein needed for translation of capped messages, and protein 3CD is transported to the nucleus where it blocks cellular transcription. These types of disruptions of cellular metabolism block antiviral responses by the cell and free the translation system to produce predominantly viral gene products. Thus, picornavirus replication can be very efficient, producing new virions after an eclipse period of less than 3 hours at yields of up to 106 virions per cell. Picornaviruses do not have a defined mechanism of cellular exit, and large paracrystalline arrays accumulate in the infected cells (Fig. 26.3).