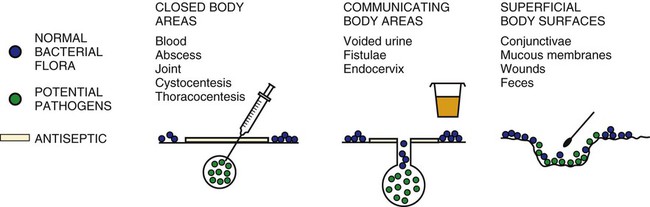
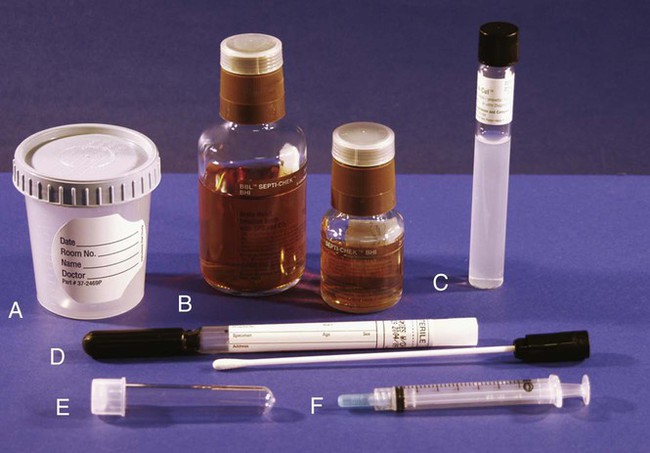
Major technologic advances have changed the way microbiology laboratories function. Bacterial identification is now achieved largely through the use of miniaturized packaged systems, many of which are automated. Antimicrobial susceptibility testing is commonly accomplished with a commercial microdilution system or some form of instrumentation. Immunodiagnostic methods for antigen and antibody detection have benefited greatly from radioimmunoassay, direct and indirect fluorescent antibody (FA) testing, enzyme-linked immunosorbent assay, latex agglutination, and immunoblotting, as well as the development of monoclonal antibody technology. Nucleic acid technology is leading the clinical microbiology laboratory into a new era of molecular diagnostics for detecting and identifying microorganisms. Cost constraints for new technology may limit the scope of services that can reasonably be provided by any one laboratory. Therefore, it is unreasonable to expect equal diagnostic capabilities from all microbiology laboratories for all microorganisms. With increasing frequency, specimens must be sent to reference laboratories that perform specialized diagnostic tests. Web Appendix 5 lists commercial, state, and federal diagnostic laboratories that provide specific microbiologic testing for infectious diseases of dogs and cats. Direct microscopic examination of exudates or infected body fluids is the single most important and cost-effective laboratory procedure that can be done for diagnosis and management of bacterial infections. Evaluations can be done in various clinical settings to obtain immediate information on the number, morphologic characteristics, and Gram-staining properties of microorganisms (Table 29-1) and the host cellular response. Purulent inflammatory responses are most suggestive of bacterial infection. Microscopic examination also gives an indication of the suitability of the specimen for culture, the likelihood of the presence of infection, the likely pathogens, and the predominant organisms in a mixed infection. This information may be used as a basis for interpretation of the significance of subsequent culture results. TABLE 29-1 Appearance of Clinical Material in Gram-Stained Preparations Certain infectious diseases may be more efficiently diagnosed by direct detection of antigens, nucleic acids, or toxins. For diagnosis of some clostridial diseases, such as botulism (see Chapter 40), tetanus (see Chapter 41), and enterocolitis (see Chapter 37), it is imperative to identify the toxin rather than rely solely on isolation of the bacteria. Direct or indirect FA stains are extremely helpful for the rapid presumptive diagnosis of infection with some bacteria. Because Yersinia pestis (see Chapter 45) and Francisella tularensis (see Chapter 46) are hazardous to laboratory personnel, antigen detection by direct FA staining of exudate or tissue is the best tool for their rapid and specific identification. DNA probes and nucleic acid amplification techniques are useful for identification of microorganisms for which culture and serologic methods are difficult, extremely expensive, or unavailable. Specimens must be collected from the actual site of infection with a minimum of contamination from adjacent tissues, organs, or secretions (Fig. 29-1). They must then be transported to the laboratory without further contamination or change in the relative numbers of bacteria. Some specimens are likely to become contaminated with indigenous flora during the collection process. Great care must be taken to use aseptic technique when collecting specimens to reduce the amount and likelihood of contamination. Some specimens are simply inaccessible except by aspiration or biopsy of deeper tissues. For all specimens obtained by biopsy or fine-needle aspiration, skin decontamination should be performed as it is before surgery. Specimens should be collected as early in the course of the disease process as possible. As the disease progresses and necrosis of tissue occurs, some microorganisms may die or be overgrown by other bacteria. Appropriate collection devices and specimen transport systems are needed to ensure survival of the microorganism without overgrowth and for optimal isolation and identification. Various containers are commercially available, ranging from simple swab and plastic tube combinations to complicated specimen collection devices (Fig. 29-2). Swabs must be made of noninhibitory materials and transported in a sterile container. Many bacteria are susceptible to desiccation during transport. Swabs are only acceptable for transport of specimens when provided with a humidified transporting chamber or placed in transport medium. Several swab-transport medium systems exist and are often available from the clinical laboratory. Tissue, exudate, feces, or fluid should be submitted in an appropriate closed (leak-proof), sterile container. Plastic screw-cap cups and blood collection tubes that do not contain anticoagulants or preservatives are examples of recommended specimen containers. Each tissue or specimen must be placed in a separate container. Direct aspiration into a syringe is often a convenient and satisfactory means for of tissue and fluid collection. However, the needle must be removed for transport to avoid injuries, and the syringe should be capped. Various types of media may be used for transporting specimens to the laboratory. Transport media such as Stuart’s, Cary-Blair, Amies, and anaerobic devices (frequently available from the microbiology laboratory) are buffered nonnutritive formulations that preserve the viability of fastidious organisms in the specimen as well as minimize overgrowth by rapidly growing bacteria that may be present. The ordinary nutrient type of broth such as that in blood culture bottles may be used only when swabs or aspirates are collected from normally sterile body sites (e.g., CSF, synovia) and when great care has been taken to avoid any contamination. Special anaerobic transport devices must be used to prevent exposure of obligate anaerobes to lethal concentrations of oxygen (see Chapter 39). Because reduced oxygen is not lethal for the aerobes and facultative anaerobes, they can be transported in the same anaerobic transport devices. Tissue in formalin, dry swabs, and urine collected several hours earlier and not refrigerated are examples of specimens that are unsuitable for bacterial culture. Successful isolation of microorganisms from blood requires an understanding of the intermittence and low order of magnitude of most bacteremias (see Blood Culture, Chapter 86, for the appropriate technique). Several specimens must be collected for culture over a period of time. If possible, the first blood culture specimen should be drawn at the onset of fever. Another suggestion is to take three or four cultures within 1.5 to 3 hours; if more than one culture yields the same organism, it is probably significant.
Laboratory Diagnosis of Bacterial Infections
Diagnostic Methods
Direct Microscopic Examination
Component
Staining Reaction
Gram-positive bacteria
Organisms retain the crystal-violet iodine complex and appear dark blue or purple.
Gram-negative bacteria
Organisms lose the primary complex, take up the secondary dye safranin, and appear red.
Fungi (yeasts)
Organisms usually appear gram positive.
Inflammatory cells
Organisms appear gram negative. Epithelial cells may appear gram positive, gram negative, or both depending on the thickness of the smear.
Backgrounds
Organisms usually appear gram negative but may appear gram positive. Fibrin, mucus, and erythrocytes often stain gram negative.
Isolation and Identification
Specimen Collection
Collection Devices and Transport
Blood
![]()
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree


Laboratory Diagnosis of Bacterial Infections