CHAPTER 44 Genetics of Feline Diseases and Traits
The inherited diseases and traits currently described in the cat are more likely to be identified in a specific breed than in random-bred cats. More than 88 million owned cats are estimated in the United States.2 Only a small percentage of this cat population is represented by cats of a specific breed, perhaps at most 15%.63 Many traits that can also be considered diseases are actually the hallmarks or identifiers of some cat breeds. Inbreeding does not cause mutations to occur at a higher rate within a breed, although inbreeding, popular sire effects, and population bottlenecks will allow rare mutations to occur more frequently within the population. Usually, recessive mutations, which can go unrecognized for many generations because they require the causative mutation to be present on both chromosomes, are the traits and diseases that tend to suddenly appear in inbred populations. The higher likelihood of an undesired trait appearing in a breed population gives the impression that breeds are unhealthier than random-bred populations. Fortuitous and deleterious mutations occur at the same rate in cats, regardless of whether the cat is pedigreed or random bred. Fancy-breed cats are more likely to have a higher standard of health care and are more likely to be closely observed than the random-bred alley cat or housecat. Thus ascertainment bias contributes to the identification of deleterious traits that are presented to the veterinarian. However, many mutations have been found in random-bred domestic shorthairs. This chapter reviews the known inherited diseases of the cat from the genetic point of view. Additional details regarding the diagnosis and treatment of the specific diseases can be found in other chapters in this volume. A list of common genetic terms can be found in Box 44-1. Diseases or conditions that are caused by genetic abnormalities cannot be cured, but the associated health problems may be manageable. An overall goal for identifying the genetic mutations for genetic conditions is the correction of the defect by way of gene or stem cell therapies or better management by way of designer drug therapies. Genetic testing is currently an effective preventive medicine because proper breeding can prevent the birth of diseased individuals; moreover, genetic testing may lead to the potential ultimate cure.
BOX 44-1 Glossary of Genetic Terms
Allele: Alternative form of a gene. One of the different forms of a gene that can exist at a single locus.
Base pair: The nucleotides that constitute a strand of DNA are also known as bases. A base pair implies the nucleotide from one strand of DNA and the pair with which it forms hydrogen bonds on the second strand of DNA, such as cytosine binding with guanine or adenine binding with thymine. The two strands of DNA are bound together by the hydrogen bonds of base pairs to form the double helix of DNA.
Common ancestry: The state of two individuals when they are blood relatives. When two parents have a common ancestor, their offspring will be inbred.
DNA (deoxyribonucleic acid): An antiparallel double helix of nucleotides (having deoxyribose as their sugars) linked by phosphodiester (sugar–phosphate) bonds to adjacent nucleotides in the same chain and by hydrogen bonds to complementary nucleotides in the opposite chain. The fundamental substance of which genes are composed.
Dominant allele: An allele that expresses its phenotypic effect even when heterozygous with a recessive allele; thus if A is dominant over a, then AA and Aa have the same phenotype.
Exon: A region of a gene that is present in the final functional transcript (mRNA) from that gene. Any non-intron section of the coding sequence of a gene; together the exons constitute the mRNA and are translated into protein.
Gene: Segregating and heritable determinant of the phenotype. The fundamental physical and functional unit of heredity, which carries information from one generation to the next. A segment of DNA, composed of a transcribed region and regulatory sequences that make possible transcription.
Gene interaction: The collaboration of several different genes in the production of one phenotypic character (or related group of characters).
Gene locus: The specific place on a chromosome where a gene is located.
Gene mutation: Mutation (point or larger change) that results from changes within the structure of a gene.
Genetic polymorphism: The occurrence together in the same population of more than one allele or genetic marker at the same locus with the least frequent allele or marker occurring more frequently than can be accounted for by mutation alone.
Genetic variance: Phenotypic variance resulting from the presence of different genotypes in the population.
Genetics: (1) The study of genes through their variation. (2) The study of inheritance.
Genome: The entire complement of genetic material in a chromosome set. The entire genetic complement of a prokaryote, virus, mitochondrion, or chloroplast or the haploid nuclear genetic complement of a eukaryotic species.
Genotype: The specific allelic composition of a cell, either of the entire cell or more commonly for a certain gene or a set of genes. The genes that an organism possesses.
Heritability: A measure of the degree to which the variance in the distribution of a phenotype is due to genetic causes. In the broad sense, it is measured by the total genetic variance divided by the total phenotypic variance. In the narrow sense, it is measured by the genetic variance due to additive genes divided by the total phenotypic variance.
Heterozygosity: A measure of the genetic variation in a population; with respect to one locus, stated as the frequency of heterozygotes for that locus.
Heterozygote: An individual having a heterozygous gene pair. A diploid or polyploid with different alleles at a particular locus.
Inbreeding: The mating of genetically related individuals. Mating between relatives.
Inbreeding depression: A depression of vigor or yield due to inbreeding.
Incomplete dominance: The situation in which both alleles of a heterozygote influence the phenotype. The phenotype is usually intermediate between the two homozygous phenotypes. The situation in which a heterozygote shows a phenotype somewhere (but not exactly halfway) intermediate between the corresponding homozygote phenotypes. (Exact intermediacy is not dominance.) See also dominance, co-dominance, and recessivity.
Intron (intervening sequence): A DNA segment of largely unknown function within a gene that specifically interrupts the coding (exon) sequences of that gene. Introns are transcribed as part of the normal gene primary transcript, but intron sequences are not found in the functional mRNA. Intron sequences are removed from the primary transcript by a splicing mechanism.
Locus (plural, loci): The position of a gene, DNA marker, or genetic marker on a chromosome.
Mosaic: A chimera; a tissue containing two or more genetically distinct cell types, or an individual composed of such tissues. Individual made up of two or more genetically distinct cell lines.
Mutant allele: An allele differing from the allele found in the standard or wild-type organism.
Mutation: (1) The process producing a gene or a chromosome differing from the wild-type. (2) The gene or chromosome that results from such a process.
Mutation rate: The number of mutation events per gene per unit of time (e.g., per cell generation). The proportion of mutations per cell division in bacteria or single-celled organisms or the proportion of mutations per gamete in higher organisms.
Nucleotide: One of four bases that constitute DNA, including adenine, guanine, cytosine, and thymine.
Pedigree: A family tree drawn with standard genetic symbols, showing inheritance patterns for specific phenotypic characters. A representation of the ancestry of an individual or family; a family tree.
Penetrance: The proportion of individuals with a specific genotype who manifest that genotype at the phenotype level.
Phenocopy: A phenotype that is not genetically controlled but looks like a genetically controlled phenotype. An environmentally induced phenotype that resembles the phenotype produced by a mutation.
Phenotype: (1) The form taken by some character (or group of characters) in a specific individual. (2) The detectable outward manifestations of a specific genotype. (3) The observable attributes of an organism.
Pleiotropy: The phenomenon whereby a single mutation affects several apparently unrelated aspects of the phenotype.
Point mutation: A mutation that can be mapped to one specific site within a locus. A small mutation that consists of the replacement (transition or transversion), addition, or deletion (frameshift) of one base.
Popular sire: A specific male in a population whose genetics becomes overrepresented in the next generation as a result of excessive breeding. Popular sires are determined by some quality desired in the population and generally represent high-winning individuals for a breed in competition. A undesired recessive trait can quickly spread in a population as a result of the popular sire effect.
Recessive: (1) An allele that is not expressed in the heterozygous condition. (2) The phenotype of the homozygote of a recessive allele.
Selection: Genetic breeding methods start with selecting particular desirable phenotypes as parents for the next generation.
Sex chromosome: A chromosome whose presence or absence is correlated with the sex of the bearer; a chromosome that plays a role in sex determination. Heteromorphic (different-shaped [e.g., X and Y]) chromosomes whose distribution in a zygote determines the sex of the organism.
Sex linked: The inheritance pattern of loci located on the sex chromosomes (usually the X chromosome in XY species); also refers to the loci themselves.
Silent mutation: Mutation in which the function of the protein product of the gene is unaltered.
X chromosome inactivation: In female mammalian embryos, the early random inactivation of the genes on one of the X chromosomes, leading to mosaicism for functions coded by heterozygous X-linked genes.
Hallmarks of Genetic Diseases
The following are six common hallmarks for inherited diseases:
Only advanced age of parents at birth has not been shown to have an effect in feline inherited diseases to date. Examples of parental age effects in humans include older or very young mothers having a higher frequency of children with trisomy 21 (Down syndrome)77 and certain types of dwarfism being associated with advanced paternal age.78
Two examples of diseases that present as sporadic and inherited forms are kidney cysts13 and lymphosarcoma.63 Each of the five characteristics that define genetic diseases can help differentiate cats with polycystic kidney disease (PKD) from cats with sporadic kidney cysts. Kidney cysts can occur in any cat, but not all cystic presentations are indicative of PKD. PKD can sometimes be detected by ultrasound as early as 6 to 8 weeks of age, consistently by 10 months of age.26 Both kidneys are generally affected, and multiple cysts are generally present (see Figures 32-4 and 32-5). The cysts are not similar in size but are similar in etiology. PKD is rampant in Persian cats and therefore must be considered a health concern in related breeds, such as Exotic Shorthairs and Himalayans. Surprisingly, this genetic problem has a very high frequency in one of the oldest and largest cat breeds, which is not a small or closed population, but the early onset, the bilateral presentation, and the high prevalence in a breed clearly demarcate this condition as a heritable problem. An older, random-bred cat with one or a few cysts in one kidney would not be a candidate for heritable PKD and genetic testing.
Lymphosarcoma is also common in cats but generally found in older cats and cats that have been infected with feline leukemia virus (FeLV).33 Mediastinal lymphosarcoma has been specifically identified in Oriental Shorthairs that are FeLV negative and generally younger than 2 years of age,31,32 although a genetic cause has not yet been identified. Other related breeds, such as Siamese, Colorpoint Shorthair, and the longhaired varieties of Siamese, have an increased prevalence of this type of lymphoma. The tumors respond well to chemotherapy, but reoccurrence is high, and the disease generally carries a very poor prognosis. This disease is found in a closed, inbred population; it has an early onset and a generally uniform presentation; and the tumor is found in areas not common to older-onset forms of lymphosarcoma. These hallmarks strongly suggest that mediastinal lymphoma is a heritable condition in the Oriental cats.
Non-genetic components (e.g., toxins, infections, infestations, sporadic damage and changes to the DNA, and environmental influences such as diet, exercise, and social surroundings) can produce a phenotype that looks just like an inherited characteristic or disease; this is termed a phenocopy. Detailed examinations of cats with heart murmurs may reveal different presentations of heart disease, one that may be genetic and one that may be environmentally induced, such as by insufficient dietary taurine resulting in dilated cardiomyopathy.82 Some diseases may present differently in different tissues, which is termed pleiotropic effects of the same gene. For example, some completely white cats show only the white coat color; others have blue eyes or one blue and one green eye, and some may be deaf.4,10,99 The variations in eye color and hearing are pleiotropic effects of the White gene for cats. Genetic testing can help the clinician rule out the common and environmental causes of clinical presentations as opposed to a condition caused by a heritable defect in the cat’s DNA.
Simple Genetic Traits
Genetic Traits with Known Mutations
The comparative genetics approach is also often termed a “candidate-gene” approach. Discovered in the early and mid-1990s, the first mutations identified in cats were for lipid and lysosomal storage diseases100,102 because these diseases have well-defined phenotypes and known genes with mutations that were as found in humans (see reviews by Banks and Chamberlain,6 and Valayannopoulos et al97). Most of the common diseases, coat colors, and coat types have been deciphered in the cat following the same candidate-gene approach, by finding a replicate trait in another species, usually mice, and checking the same gene for causative mutations.
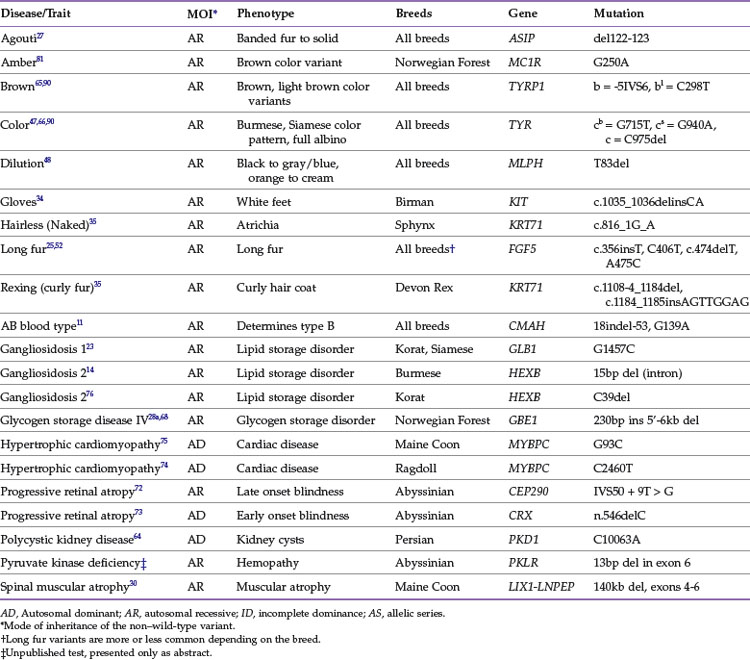
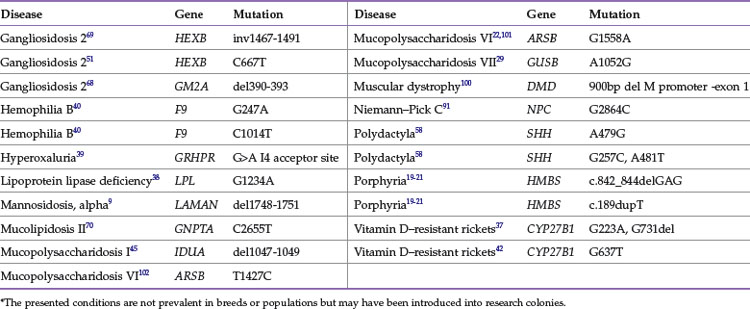
Once a mutation is known, a genetic-based test can be established. DNA testing for domestic cat diseases and appearance traits is a rapidly growing asset for the veterinary community. Approximately 33 genes contain approximately 50 mutations that cause feline health problems or alterations in the cat’s appearance (Tables 44-1 and 44-2). To date, other than the muscular dystrophy mutation,100 all mutations in the cat are autosomal, not found on the X or Y chromosomes. A variety of commercial laboratories can now perform feline genetic diagnostics, allowing both the veterinary clinician and the private owner to obtain DNA test results. DNA is easily obtained from a cat by buccal swab using a standard cotton bud or cytologic brush, after which DNA samples can be sent to any laboratory in the world because the DNA is stable at room temperature. The DNA test results identify carriers of the traits, predict the incidence of traits in breeding programs, and influence medical prognoses and treatments. Once a genetic test proves that an animal has a genetic trait, preventive therapies and dietary restrictions could be implemented to slow or prevent the onset and progression of the associated disease. Thus genetic testing should not be viewed as an alternative to veterinary care but instead as a tool for veterinary care, part of a cat’s overall health management plan.
Phenotypic Mutations of the Domestic Cat
Domestic cats have been selected to produce breeds mainly on the basis of aesthetic qualities, especially coloration, fur length and type, and some morphologic types such as folded or curled ears. Most of the genes controlling these traits are simple, and many of the causative mutations have been identified. Tables 44-1 and 44-3 present the common genes and loci that affect feline phenotypic traits. A trait is initially given a locus name, such as Brown, before the actual gene has been identified. The locus name usually is a descriptor of the trait, although the location in the genome for the locus will not be initially known; however, the mode of inheritance of the different alleles is usually determined. The alleles are given single- or two-letter designations, lower case implying a recessive allele. Once the gene is identified, such as tyrosinase-related protein 1 (TRYP1) for Brown, the mutations are written to describe the genetic alteration within the gene, and the gene will be designated with the alleles, such as TRYP1b for the brown allele.
TABLE 44-3 Simple Phenotypic Traits and Diseases of the Cat and Its Breeds: Mutations Are Unidentified
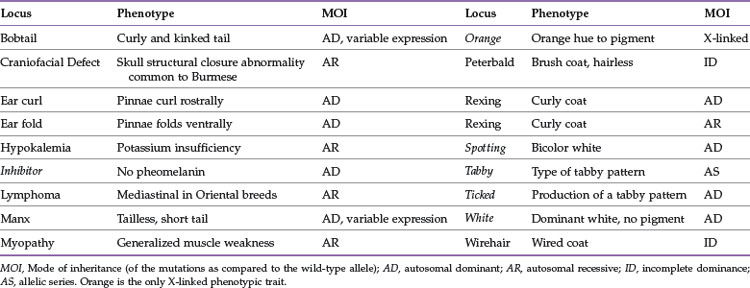
Several of the cat coat color loci have only one mutant allele, such as Agouti, Dilute, and Extension. Because the phenotypic mutations are of value to cat breeders for managing their breeding programs, the phenotypic mutations generally have genetic tests readily available from commercial services. Because most of the mutations for the aesthetic traits are recessive, the mutant alleles must be present in both copies for the effect to be visible, and cats can carry the mutation without detection. Recessive mutations tend to be found in the genes that produce the enzymes of biological pathways. Thus most of the coat color mutations are recessive because they are usually part of disrupting the pigment production pathways. Many genes that affect pathways also tend to have more than one mutation that cause different effects; this is termed an allelic series. The locus for brown color variants, Brown, has two mutations in the causative gene, TRYP1. The wild-type allele, B, which causes normal black pigment, is dominant to the brown allele, b, which causes a reduced amount of black pigment, producing a more brownish hue to the fur. The brown allele, b, is considered dominant to light brown, whereas bl imparts a cinnamon-color or reddish effect on the fur. The allelic series is written as: B > b > bl to indicate the dominance of one allele over the other. In the case of the Color locus, C, which is also an allelic series, the sepia coloration, cbcb, which is fixed in the Burmese (Figure 44-1) and Singapura (Figure 44-2) breeds, is co-dominantly expressed with the Siamese points, cscs, producing an additive affect. Thus compound heterozygous cats, cbcs, have an intermediate coloration compared with that of the Burmese and the Siamese; this is usually referred to as a mink Tonkinese (Figure 44-3). Complete albinos, which have an additional allele at the Color locus, have been identified. The locus is controlled by the gene tyrosinase; TYR and the allelic series is written as C > cb = cs > c.
The coat color mutations are common to all cats and are effective for genetic typing in all breeds and populations. However, even though long fur is common in pedigreed and random-bred cats, long fur is an exception because four different mutations in the gene fibroblast growth factor 5 (FGF5) can cause a cat to have long fur.25,52 One mutation is common to almost all breeds and populations, which suggests that this mutation is the most ancient and present before breeds developed, but the other long fur mutations are more specific to particular breeds.3 Some cats can have long fur because of two different mutations in the gene FGF5. These cats would be considered compound heterozygotes. Thus all four mutations must be genotyped to determine whether a cat carries a mutation for long fur.
Rexing is an interesting set of mutations for the domestic cat. At least six different types of rexoid mutations have been noted in the cat, including Cornish Rex (Figure 44-4) and Devon Rex (Figure 44-5), as well as several unpublished varieties, including American Wirehair (Figure 44-6), Selkirk Rex (Figure 44-7), LaPerm (Figure 44-8), and Tennessee Rex. The Devon Rex and Cornish Rex had been known to be caused by different genes after cross-breeding experiments and genetic studies.85 The genetic studies have ruled out the Devon Rex gene, keratin 71 (KRT71), as causative for the other rexoid cats.35 The Selkirk and LaPerm have dominant mutations, Devon and Cornish are caused by recessive mutations, and the wirehair appears to be a dominant mutation with variable expression and even incomplete penetrance, wherein sometimes cats thought to have the mutation are not wirehaired. Sometimes two traits are not initially known to be caused by the same gene, such as hairlessness of the Sphynx and rexing in the Devon Rex.35,86 Thus the hairlessness of the Sphynx (Figure 44-9) has had the locus name Hairless with alleles Hr and hr, whereas Devon Rex has had the locus name of Rex with alleles Re and re defining Devon Rex as a separate locus from Rex, with alleles R and r for the Cornish Rex. Both Sphynx and Devon Rex have been proved to be caused by mutations in KRT71.35
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree