Coronaviridae
Abstract
This chapter describes the properties of coronaviruses and features of the diseases they cause in animals.
Keywords
Coronaviridae; Torovirus; SARS; MERS; coronavirus
The family Coronaviridae is included with the families Arteriviridae, Roniviridae, and Mesoniviridae in the order Nidovirales; viruses in these four families share a distinctive replication strategy. The family Coronaviridae is comprised of two subfamilies. One, the subfamily Coronavirinae, contains a substantial number of pathogens of mammals and birds that individually cause a remarkable variety of diseases, including pneumonia, reproductive disease, enteritis, polyserositis, sialodacryoadenitis, hepatitis, encephalomyelitis, nephritis, and various other disorders (Table 24.1). Coronavirus and coronavirus-like infections have been described in swine, cattle, horses, camels, cats, dogs, rodents, birds, bats, rabbits, ferrets, mink, and various wildlife species, although many coronavirus infections are subclinical. In humans, coronaviruses are included in the spectrum of viruses that cause the common cold as well as more severe respiratory disease—specifically, severe acute respiratory syndrome (SARS) and Middle East respiratory syndrome (MERS), which are both zoonoses. The second subfamily, Torovirinae, contains pathogens of both terrestrial and aquatic animals. The genus Torovirus includes the type species, equine torovirus (Berne virus), which was first isolated from a horse with diarrhea, and Breda virus, which was first isolated from neonatal calves with diarrhea. Berne virus neutralizing antibodies have been detected in sera of sheep, goats, rabbits, and mice, and torovirus-like particles have also been observed by electron microscopy in feces of swine, cats, turkeys, and humans. White bream virus from fish is the type species of the genus Bafinivirus.
Table 24.1
| Virus | Disease/Symptoms | Transmission | Prevention/Control |
| Subfamily Coronavirinae | |||
| Genus Alphacoronavirus | |||
| Feline coronavirus (Feline enteric coronavirus; Feline infectious peritonitis virus) | Feline enteric coronavirus: mild gastroenteritis and diarrhea | Feline enteric coronavirus: direct contact; fecal–oral route from maternal shedding | Interruption of transmission cycle, quarantine, high-level hygiene |
| Feline infectious peritonitis virus: peritonitis, pneumonia, CNS signs etc. | Feline infectious peritonitis virus: blood, body fluids | ||
| Canine coronavirus | Mild gastroenteritis and diarrhea | Fecal–oral route | Inactivated vaccine |
| Possible severe enteritis and systemic signs (leucopenia) | |||
| Transmissible gastroenteritis (TGE) virus of swine | Gastroenteritis. Watery diarrhea, vomiting, dehydration | Fecal–oral route | Oral attenuated vaccine to pregnant sows. Good sanitation |
| Porcine respiratory coronavirus | Mild respiratory disease or subclinical | Aerosol | No vaccine available |
| Nasal swabs; trachea, lung sections | |||
| Porcine epidemic diarrhea virus (PEDv) | Gastroenteritis. Watery diarrhea, vomiting, dehydration | Fecal–oral route | Inactivated and oral live-attenuated virus vaccine to pregnant sows. Good sanitation |
| Genus Betacoronavirus | |||
| Group A | |||
| Porcine hemagglutinating encephalomyelitis virus | Vomiting, wasting disease, encephalomyelitis. Anorexia, hyperesthesia, muscle tremors, emaciation | Aerosols, oronasal secretions | Good husbandry, maintain immune sows |
| No vaccine available | |||
| Mouse hepatitis virus | Enteritis, hepatitis, demyelinating encephalomyelitis | Introduction of virus into a naïve colony: aerosols and direct contact | Depopulation. Preventive quarantine |
| Aerosols | |||
| Rat sialodacryoadenitis virus | Rhinitis, epiphora, pneumonia | ||
| Bovine coronavirus | Gastroenteritis with profuse or bloody diarrhea, dehydration, decreased milk, or respiratory disease | Fecal–oral route, aerosols, respiratory droplets | Maternal immunization: inactivated or attenuated vaccines; no vaccine for winter dysentery |
| Equine coronavirus | Gastroenteritis | Fecal–oral route | |
| Canine respiratory coronavirus | Respiratory disease | Aerosols | |
| Group B | |||
| Severe acute respiratory syndrome (SARS) coronavirus | Respiratory disease; zoonotic with bats as natural reservoir | Aerosols, oronasal secretions | No vaccines available; enhanced biosecurity for human cases |
| Group C | |||
| Middle East respiratory syndrome (MERS) coronavirus | Respiratory disease; zoonotic with camels and bats as a likey reservoir | Aerosols, oronasal secretions | No vaccines available; enhanced biosecurity for human cases |
| Genus Gammacoronavirus | |||
| Avian infectious bronchitis virus | Tracheobronchitis, nephritis | Aerosols and ingestion of food contaminated with feces | Multivalent attenuated and inactivated vaccines available. Good sanitation and testing |
| Rales, decreased egg production | |||
| Turkey coronavirus, Bluecomb virus | Enteritis | Fecal–oral route, aerosol | Inactivated virus vaccine |
| Diarrhea, depression, cyanotic skin | |||
| Genus Deltacoronavirus | |||
| Porcine deltacoronavirus | Gastroenteritis in sows and nursing pigs; low mortality in nursing pigs; clinically indistinguishable from TGE and PEDv | Fecal–oral route | No vaccine; biosecurity |
| Subfamily Torovirinae | |||
| Genus Torovirus | |||
| Breda virus (cattle) | Enteritis | Fecal–oral route | No vaccine available |
| Diarrhea, dehydration | |||
| Genus Bafinivirus | |||
| White bream virus | None observed | Assumed horizontal via water | No control method proposed |
| Fathead minnow | Hemorrhages in the eyes and skin | ||
| Necrotic nidovirus lesions in kidney, liver, and spleen | Assumed horizontal | None available | |
Properties of CORONAVIRUSES
Classification
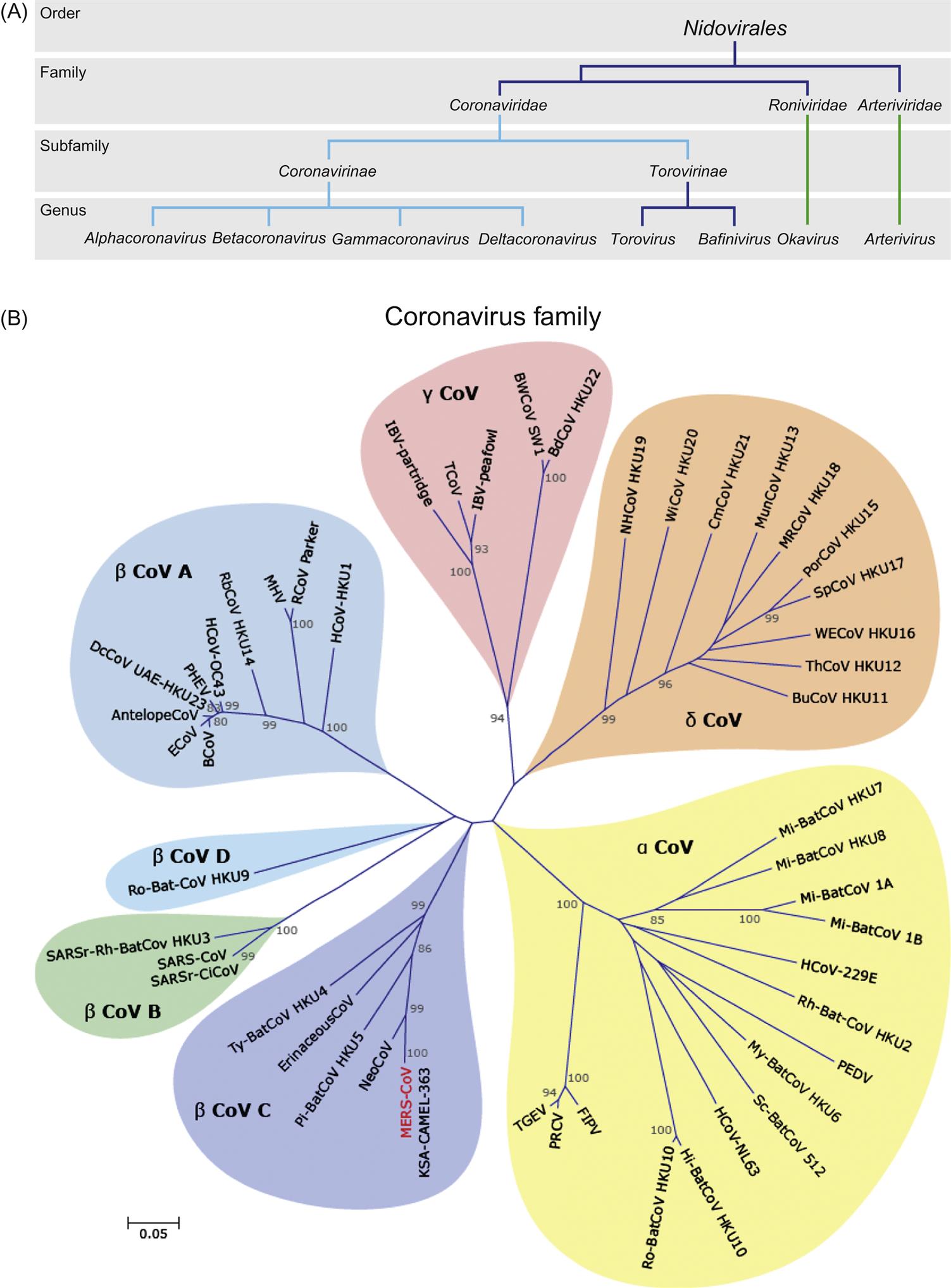
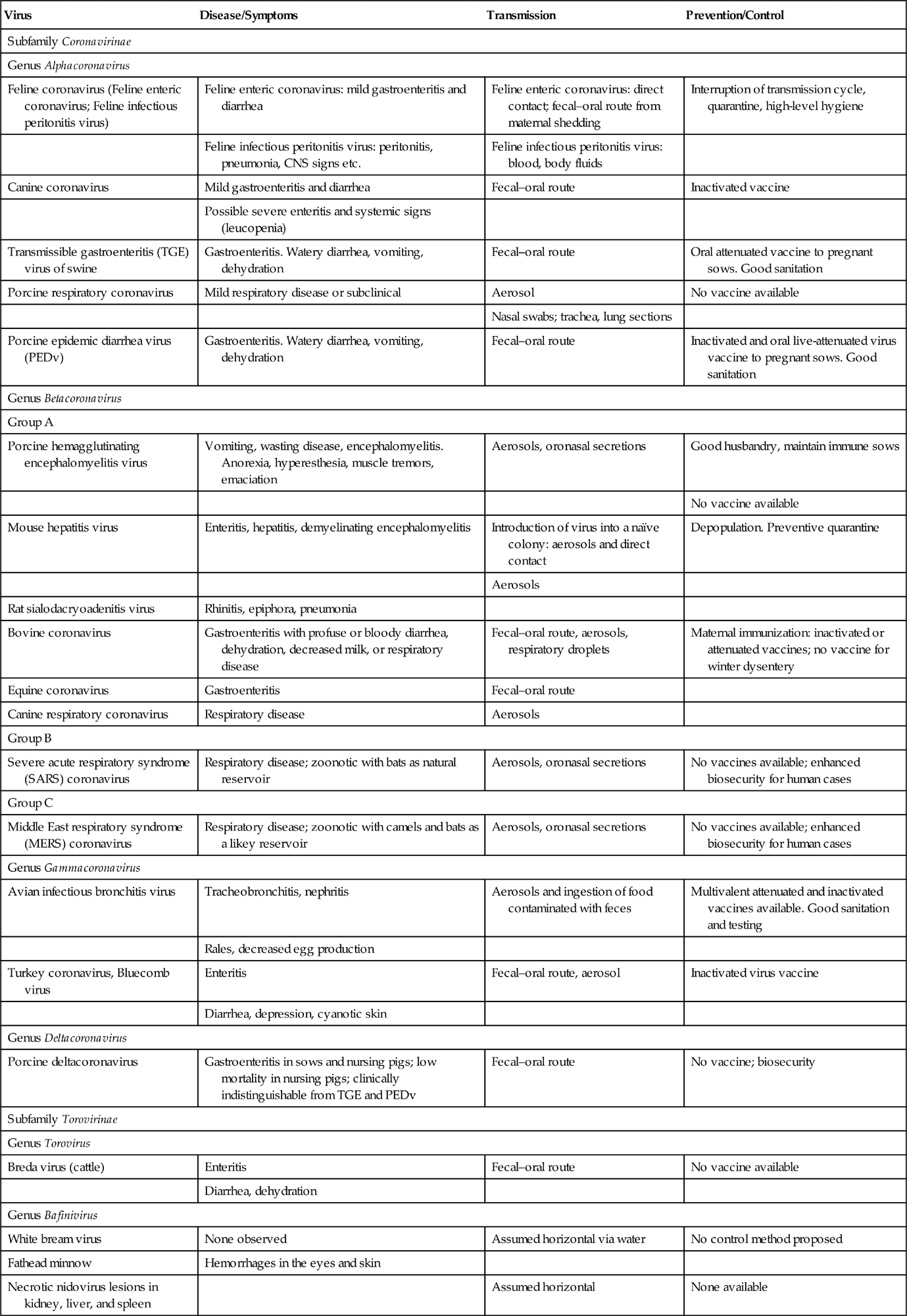
Despite profound differences in virion structure and genome size, coronaviruses, toroviruses, arteriviruses, roniviruses, and mesoniviruses exhibit remarkable similarities in their genome organization and replication strategy (Fig. 24.1; Table 24.2). In infected cells, these viruses all utilize a distinctive “nested set” transcription strategy in which the expression of genes encoding structural viral proteins is mediated via a nested set of 3′ coterminal subgenomic mRNAs. This unique strategy has been recognized by the establishment of the order Nidovirales (from the Latin nidus, nest), encompassing the family Coronaviridae, with two subfamilies (Coronavirinae and Torovirinae), and the families Arteriviridae, Roniviridae, and Mesoniviridae (Fig. 24.2A). Sequence analysis of the gene encoding portions of the viral RNA-dependent RNA polymerase (RdRp) suggests that the member viruses of the order Nidovirales probably evolved from a common ancestor. Extensive genome rearrangements through heterologous RNA recombination, along with accumulation of mutations over time, have resulted in the variations seen—that is, viruses with similar replication and transcription strategies but disparate structural features.
Table 24.2
| Proteina | Coronavirus | Torovirus | Bafinivirus | Okavirus | Arterivirus | |
| Spike glycoprotein | S | 1035–1472 | 1562–1584 | 1220 | − | − |
| Large spike glycoprotein | gp116 | − | − | − | 873c–899 | − |
| Small spike glycoprotein | gp64 | − | − | − | 539 | −− |
| Minor surface glycoprotein | GP2 | − | − | − | − | 227–249 |
| GP3 | − | − | − | − | 163–256 | |
| GP4 | − | − | − | − | 152–183 | |
| Major surface glycoprotein | GP5 | − | − | − | − | 199–278 |
| Membrane protein | M | 218–263 | 233 | 227 | − | 162–174 |
| Nucleocapsid protein | N | 349–470 | 159–167 | 161 | 144–146 | 110–128 |
| Envelope protein | E | 74–109 | − | − | − | 67–80 |
| Hemagglutinin-esterase protein | HE | 386–440b | 416–430 | − | − |
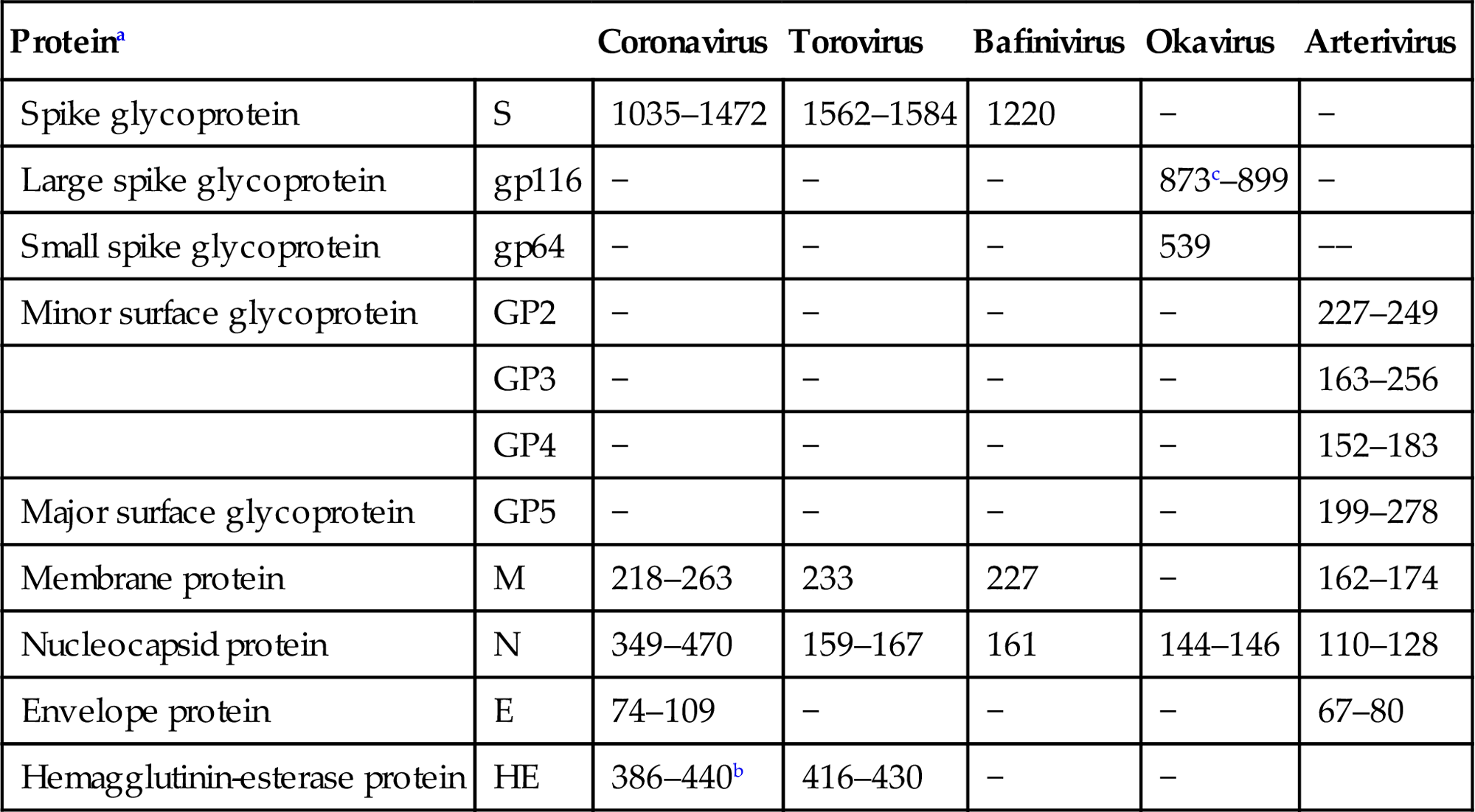
aOnly proteins typical for each lineage are listed; for some CoVs additional, virus species-specific accessory envelope proteins have been described.
bOnly found in a cluster of betacoronaviruses (“phylogroup A,” Betacoronavirus 1, Murine coronavirus, Human coronavirus HKU1).
The subfamily Coronavirinae is subdivided into four genera on the basis of genetic and serologic properties, sometimes with subgroups within these (Table 24.1; Fig. 24.2). The genus Alphacoronavirus (previously group 1 coronaviruses) includes transmissible gastroenteritis virus of swine, porcine respiratory coronavirus, porcine epidemic diarrhea virus, canine coronavirus, feline coronavirus, ferret and mink coronaviruses, the human coronaviruses 229E and HKU1, as well as many viruses found in bats. The genus Betacoronavirus (previously group 2 coronaviruses) is divided into four groups; Betacoronavirus group A includes mouse hepatitis virus, rat (sialodacryoadenitis) coronavirus, bovine and equine coronaviruses, porcine hemagglutinating encephalomyelitis virus, canine respiratory coronavirus, and other human coronaviruses. Betacoronavirus group B includes human SARS coronavirus, civet cat, raccoon dog, and horseshoe bat coronaviruses. Betacoronavirus group C includes MERS coronavirus from both humans and camels, as well as closely related bat coronaviruses, and group D currently includes only coronaviruses of bats. The genus Gammacoronavirus (previously group 3 coronaviruses) includes avian infectious bronchitis virus, turkey coronavirus, and several potential but still largely uncharacterized new species from wild birds and marine mammals, including dolphins and whales. The more recently identified genus Deltacoronavirus includes viruses from pigs and a variety of wild birds, as well as a virus from a wild Asian leopard cat. Further taxonomic subdivision of these viruses is likely in the future. It is believed currently that warm-blooded flying vertebrates are the definitive hosts for the coronavirus gene pool, with alpha- and betacoronaviruses having their origin in bats, and gamma- and delta-coronaviruses having their origin in birds.
Viruses in the Torovirinae subfamily are all apparently closely related but genetically distinct from coronaviruses; however, many toroviruses have yet to be fully characterized. There are currently two genera within the family Torovirinae, specifically, the genera Torovirus and Bafinivirus (Fig. 24.2A).
Virion Properties
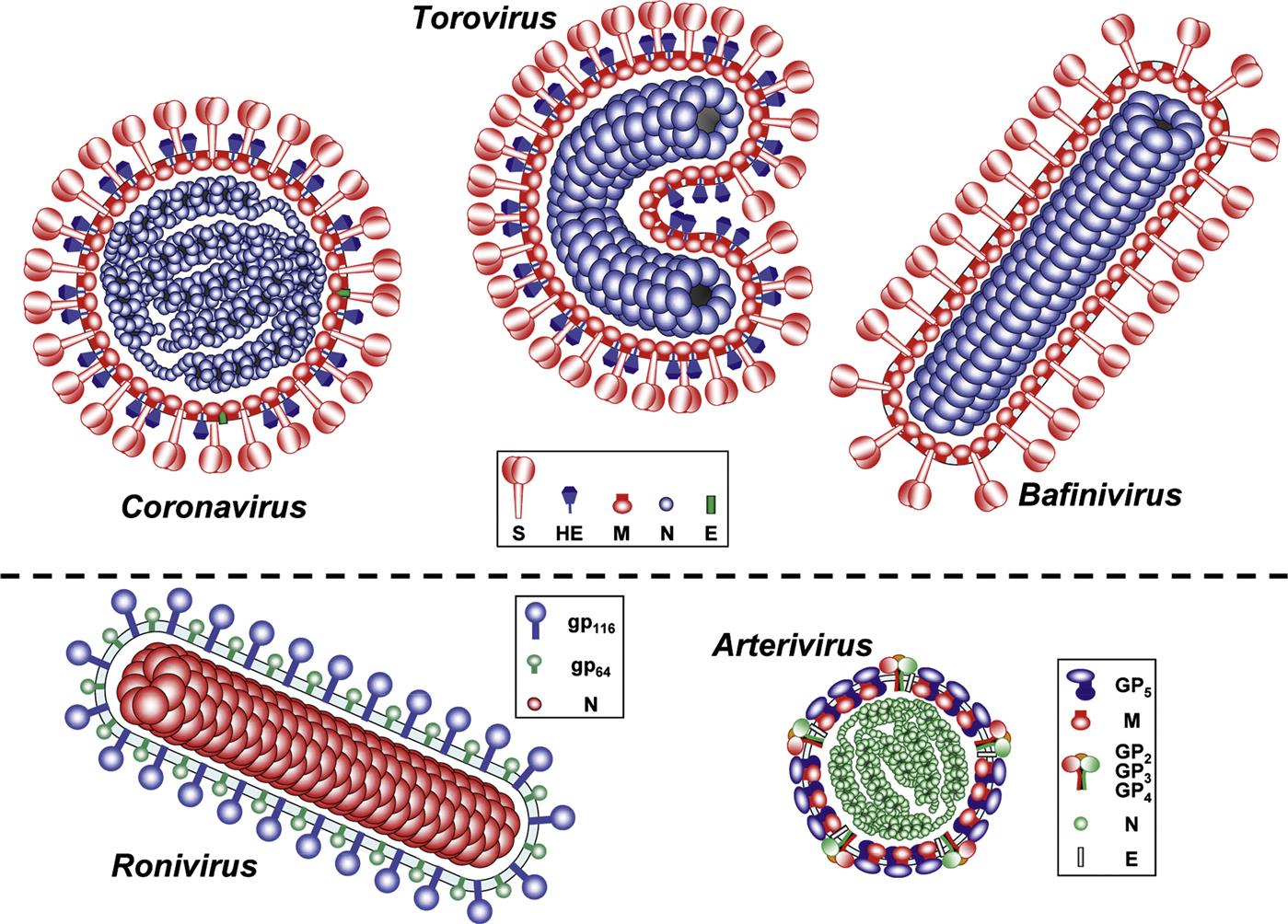
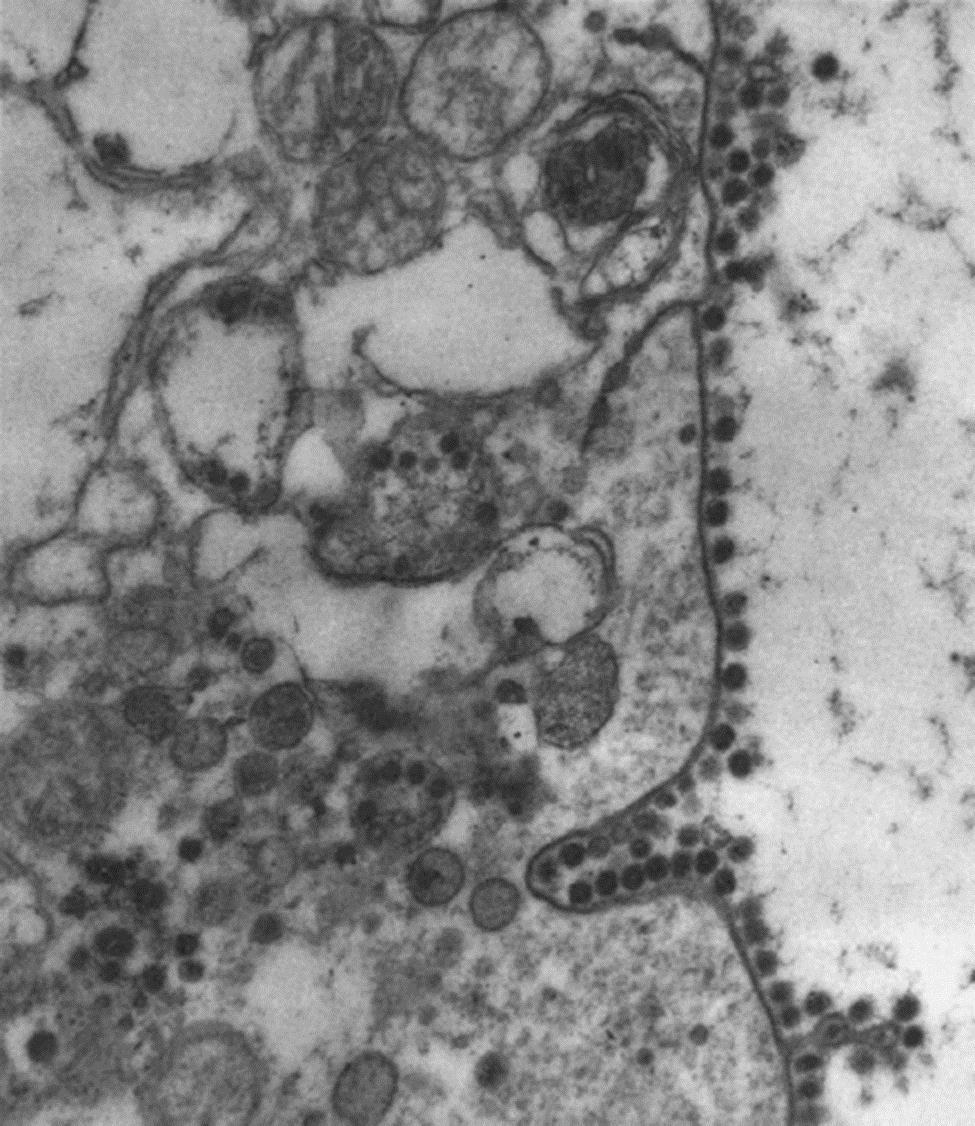
Member viruses of the family Coronaviridae are enveloped, 80–220 nm in size, pleomorphic although often spherical (coronaviruses), or 120–140 nm in size and disc, kidney, or rod-shaped (toroviruses and bafiniviruses) (Fig. 24.1). Coronaviruses have distinctive and large (20 nm long) club-shaped spikes (peplomers, composed of trimers of the spike protein). The association of the nucleocapsid (N) protein with the genomic RNA forms the helical nucleocapsid that is surrounded by an icosahedral structure composed of the viral membrane (M) protein. Some coronaviruses also have a second fringe of shorter (5 nm long) spikes (composed of the hemagglutinin-esterase (HE) protein), a particular characteristic of some betacoronaviruses. Toroviruses also have large club-shaped spikes, but the particles are more pleomorphic and have a tightly coiled tubular nucleocapsid bent into a doughnut shape. By thin-section electron microscopy, torovirus nucleocapsids appear as kidney-, disc-, or rod-shaped forms. Bafiniviruses appear as straight rods with a bacilliform morphology, which are surrounded by large peplomers.
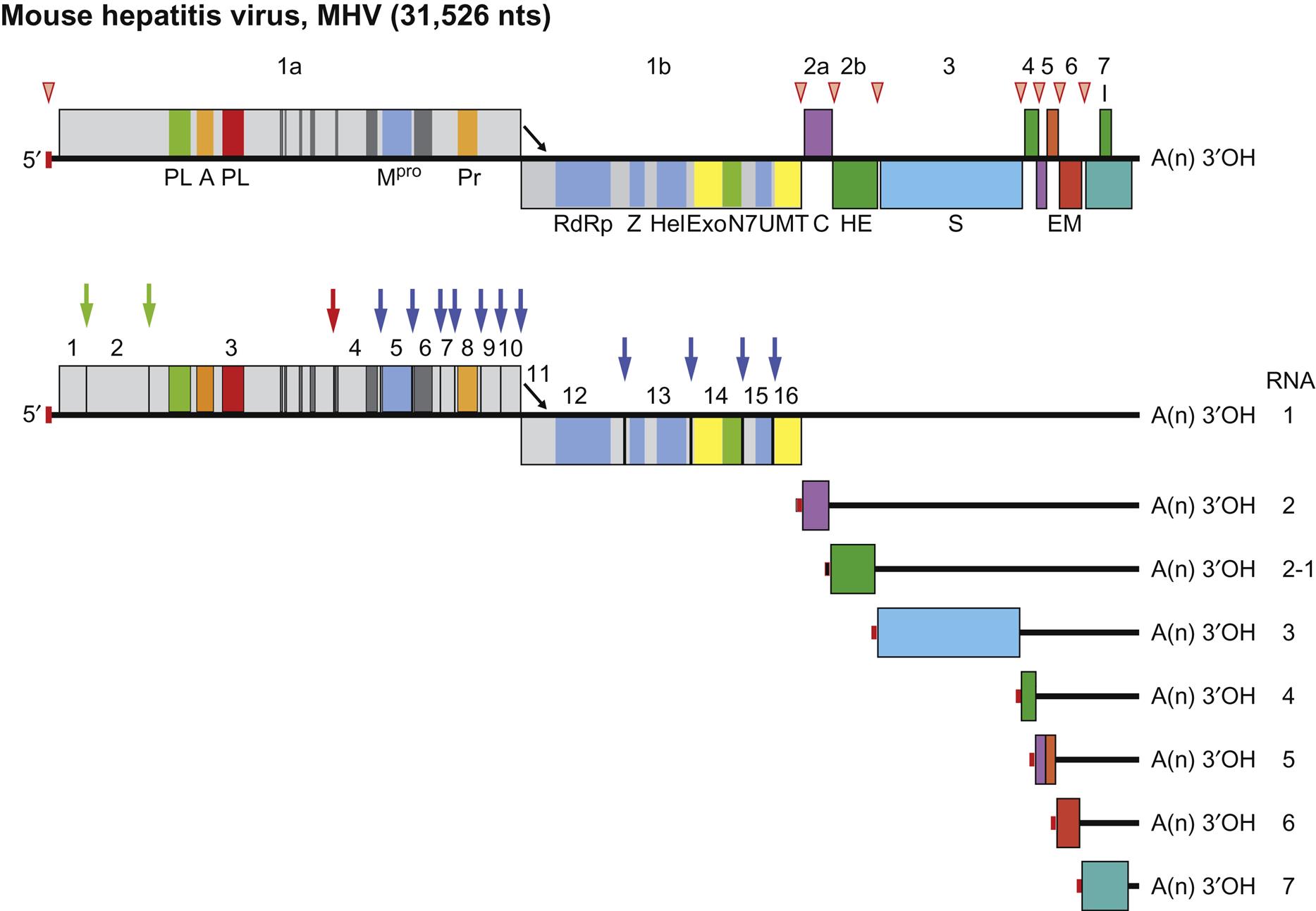
The genome of viruses in the family Coronaviridae consists of a single molecule of linear positive-sense, single-stranded RNA, 27.6–31 kb in size for coronaviruses and 25–30 kb for toroviruses, the largest known nonsegmented RNA viral genomes. The genomic RNA is 5′ capped and 3′ polyadenylated, and is infectious (Table 24.3; Fig. 24.3).
Table 24.3
Properties of Coronaviruses and Toroviruses
Virions are pleomorphic or spherical (Subfamily Coronavirinae) or disc-, kidney-, or rod-shaped (Subfamily Torovirinae); 80–220 nm (coronaviruses) or 120–140 nm (toroviruses) in diameter. Virions are enveloped, with large club-shaped spikes (peplomers)
Virions have an icosahedral core structure within which is a helical nucleocapsid (coronaviruses) or a tightly coiled tubular nucleocapsid in a doughnut (toroviruses) or bacilliform (bafiniviruses) shape
The genome consists of a single molecule of linear positive-sense, single-stranded RNA, 25–31 kb in size; the genome is 5′ capped, 3′ polyadenylated, and infectious
Coronavirus virions contain three or four structural proteins: a major spike glycoprotein (S), transmembrane glycoproteins (M and E), a nucleoprotein (N), and, in some viruses, a hemagglutinin esterase (HE). Torovirus virions contain analogous proteins, but there is no E protein. Bafiniviruses have only three structural proteins (S, M and N)
Viruses replicate in the cytoplasm; the genome is transcribed, forming a full-length complementary RNA from which is transcribed a 3′ coterminal nested set of mRNAs, only the unique sequences of which are translated
Virions are formed by budding into the endoplasmic reticulum and are released by exocytosis. Cell–cell fusion may occur
The major virion proteins of the member viruses of the subfamilies Coronavirinae and Torovirinae include a nucleocapsid protein (N, 50–60 kDa, 19 kDa for toroviruses) and several envelope proteins: (1) the spike glycoprotein trimer (S, 180–220 kDa per monomer); (2) a triple-spanning transmembrane protein (M, 23–35 kDa); (3) a minor transmembrane protein (E, 9–12 kDa), which together with the M protein is essential for coronavirus virion assembly and budding. Toroviruses lack a homolog of the coronavirus E protein, which may explain the structural differences between the coronaviruses and toroviruses (Fig. 24.1). The secondary, smaller spikes, seen in some betacoronaviruses and in toroviruses, consist of a dimer of a second class I membrane glycoprotein (65 kDa per monomer), a HE that shares 30% sequence identity with the N-terminal subunit of the HE fusion protein of influenza C virus. Sequence comparisons indicate that the HE genes of coronaviruses, toroviruses, and orthomyxoviruses were acquired by independent, nonhomologous recombination events (probably from the host cell). Although there is no sequence similarity between the torovirus proteins and their counterparts in coronaviruses, they are similar in structure and function, and are related phylogenetically. Bafiniviruses have only the S, M, and N structural proteins.
Virus neutralizing antibodies generated during natural infections are directed at the surface glycoproteins of coronaviruses and toroviruses, with the majority being conformational epitopes located at the N-terminal portion of the S protein. Cellular immune responses are principally directed toward the S and N proteins. Besides the canonical structural proteins, coronaviruses are unique among nidoviruses because their genomes encode (within differing regions) variable numbers of accessory proteins (four or five in most; eight in the SARS coronavirus) that are dispensable for in vitro virus replication, but which increase virus fitness in vivo. The accessory proteins encoded by the SARS coronavirus open reading frames 3b and 6, for example, are antagonists of innate immune responses, specifically interfering with the development of type I interferon responses (see Chapter 4: Antiviral Immunity and Virus Vaccines); the specific roles of other accessory proteins are still largely unknown. The accessory proteins have homologous versions within coronavirus groups, but lack similarity with proteins in different groups. In the betacoronaviruses, for example, the HE protein is considered an accessory protein, and mouse hepatitis virus HE-deletion mutants replicate like wild-type virus in vitro, but in mice they have an attenuated phenotype.
Virus Replication
The host spectrum/tropism of individual coronaviruses appears to be largely determined by the S protein, portions of which mediate receptor binding and virus cell fusion that occur at either the plasma membrane or within endosomes of susceptible cells. Individual coronaviruses utilize a variety of cellular proteins as receptors. Aminopeptidase N (APN or CD13) serves as a receptor for several alphacoronaviruses, including feline coronavirus, canine coronavirus, transmissible gastroenteritis virus, porcine epidemic diarrhea virus, and human coronavirus 229E. SARS coronavirus and human coronavirus NL63 utilize angiotensin converting enzyme 2 (ACE2). MERS coronavirus utilizes dipeptidyl-peptidase 4 (DPP4 or CD26). Some strains of mouse hepatitis virus utilizes carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM-1). Other betacoronaviruses utilize sialic acids as a primary receptor (eg, N-acetyl-9-O-acetyl neuraminic acid). In some cases, eg, transmissible gastroenteritis virus, the spike protein can bind to both specific and nonspecific receptors (eg, APN and sialic acids) via distinct subdomains. The functional receptor for gammacoronaviruses such as infectious bronchitis virus is undefined, although sialic acid residues may serve as nonspecific attachment factors. Many coronavirus spike proteins also interact with C-type lectins [such as liver/lymph node-specific intercellular adhesion molecule-3-grabbing integrin (L-SIGN or CD 209L) and dendritic cell-specific intercellular adhesion molecule-3-grabbing nonintegrin (DC-SIGN or CD 209)], which may serve as nonspecific attachment factors in a complex with a primary receptor. In addition to receptor binding, the activation of virus fusion via the action of host cell-specific proteases that cleave spike is likely to be a powerful means of regulating coronavirus infection and host- or tissue-tropism.
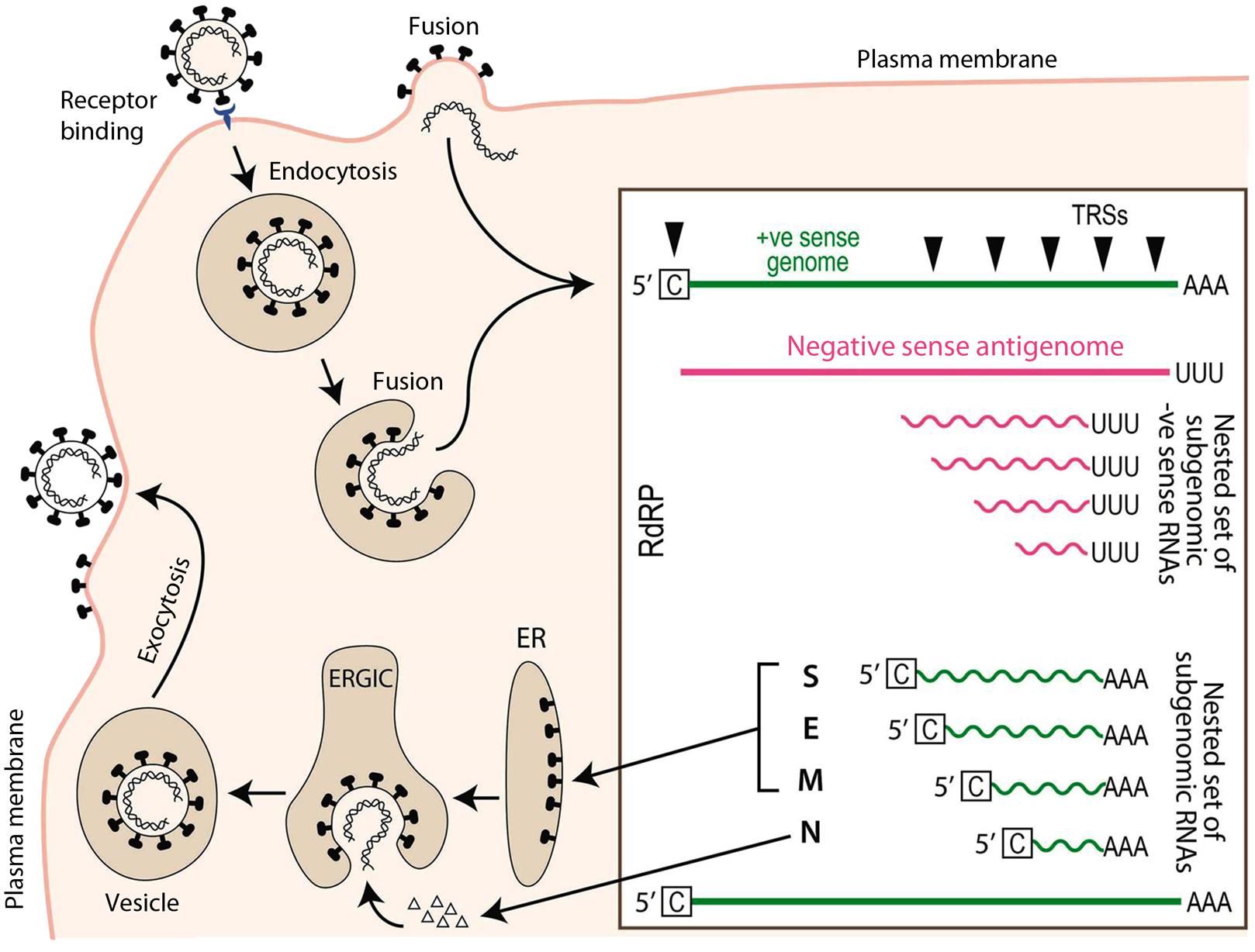
Virus replication and transcription, as for many RNA viruses, takes place within an extensive membranous network of virus-modified endoplasmic reticulum-derived vesicles. The strategy of viral replication and transcription of the coronavirus genome is complex (Figs. 24.3 and 24.4; see also chapter: Arteriviridae and Roniviridae, Fig. 25.4 which depicts the replication of another member of the Order Nidovirales). First, the viral RNA serves as messenger RNA (mRNA) for synthesis of the RNA dependent RNA polymerase (RdRp). The two large 5′-most open reading frames, ORF1a and ORF1b (some 20 kb in total size) encoding the subunits of the polymerase are translated—the larger via ribosomal frameshifting—as a single polyprotein (pp1a or pp1ab) that is then cleaved by virus-encoded proteases found within the polyprotein, resulting in the production of mature products that are termed nsp1 to nsp16 (nsp, nonstructural protein). These proteins then assemble within the network of rearranged membranes to form the active replicase–transcriptase complex, comprising the RNA polymerase (nsp12) and accessory proteins, including a 3′–5′ exonuclease that imparts some degree of “proof reading” function during replication, an unusual feature for RNA viruses. Such proof reading activity is thought to be important in maintaining the integrity of such large RNA genomes and in avoiding accumulation of excessive numbers of mutations associated with RNA polymerase infidelity.
The viral polymerase is used to synthesize full-length negative-sense (complementary) RNA by copying the genome starting at the 3′ end. The antigenome is then copied back into full-length positive-sense genomic RNA. The generation of full-length genomic RNA is done utilizing the replicase activity of the viral RNA dependent RNA polymerase. In addition, the RNA dependent RNA polymerase can also synthesize a nested set of RNAs with different sizes that are generated by a discontinuous synthesis of negative-sense RNAs. This is done using the transcriptase activity of the RNA dependent RNA polymerase. In this case, the RNA dependent RNA polymerasesynthesizes negative-sense RNA by starting to copy at the 3′ end of the genome, it then recognizes internal regulatory sequences, the transcriptional regulatory sequences (TRSs) found upstream of each open reading frame, where it pauses and translocates to the 5′ end of the genome, guided by sequence complementarity. The RNA dependent RNA polymerase then extends the nascent negative-sense RNA by copying the leader sequence found at the genome’s 5′ end. These negative-sense template RNAs, sharing both 5′ and 3′ ends, are copied into positive-sense subgenomic mRNAs which then allow expression of viral genes downstream of the replicase. The template switching employed during transcription is at the heart of the RNA recombination that is a hallmark of coronavirus replication.
In addition to the accumulation of point mutations as a result of polymerase errors (infidelity) during transcription (genetic drift), genetic recombination occurs at high frequency between the genomes of different but related coronaviruses during coinfection situations. Recombination between coronaviruses is a direct result of the discontinuous transcription strategy employed by the viral polymerase, and the presence of transcriptional regulatory sequences in the viral genome. Such recombination is likely to be an important mechanism for the generation of the genetic diversity seen with these viruses in nature, and provides a constant potential source of new viruses with novel phenotypic properties, such as host range, tissue tropism, and virulence.
Among members of the subfamily Torovirinae, transcription and replication apparently are similar to those of coronaviruses, except that there are no common 5′ leader sequences on the mRNAs of viruses in the genus Torovirus. As occurs during replication of coronaviruses, subgenomic negative-sense RNAs complementary to the nested set of mRNAs are also present in torovirus-infected cells. The fact that these subgenomic RNAs contain 5′- and 3′-terminal sequences that are identical to those of genomic RNA implies that they may function as replicons. Viruses in the genus Bafinivirus use the same transcriptional strategy as the coronaviruses and produce their replicase polyproteins from the virus genome and the three structural proteins from a nested set of 3′-coterminal subgenomic mRNAs, each having a common 5′ leader sequence identical to that of the virus genome.
The synthesis, processing, oligomerization, and transport of the several envelope glycoproteins of coronaviruses display some unusual features. For example, the envelope protein M, which in some coronaviruses contains O-linked rather than N-linked glycans, is directed exclusively to the cisternae of the endoplasmic reticulum and other pre-Golgi membranes. As a result, virions bud into the lumen of the endoplasmic reticulum–Golgi and not from the plasma membrane. Assembled virions are transported in Golgi-derived vesicles to the plasma membrane, where they are released by exocytosis (Fig. 24.5). After their release, many of the mature enveloped virions remain adherent to the outside of the cell. The spike proteins are coassociated with M at the endoplasmic reticulum–Golgi interface, but are also expressed on the cell surface where they can trigger extensive cell–cell fusion, resulting in syncytia formation.
Members of the Subfamily Coronavirinae
The subdivision of viruses included in the subfamily Coronavirinae into genera (Alpha-, Beta-, Delta-, and Gammacoronaviruses) is based largely on comparative genome sequence analyses, rather than the biological properties of individual viruses. Thus, these viruses will be grouped according to the animal species they infect, rather than their taxonomic assignment; specifically, coronaviruses of birds (infectious bronchitis, turkey coronavirus), cats, dogs and ferrets (feline enteric coronavirus, feline infectious peritonitis virus, canine coronavirus, canine respiratory coronavirus, ferret coronavirus), cattle and horses (bovine coronavirus, equine coronavirus), laboratory animals (mouse hepatitis virus, rat coronavirus (rat sialodacroadenitis coronavirus), guinea pig and rabbit coronaviruses), swine (transmissible gastroenteritis, porcine epidemic diarrhea, porcine respiratory corona virus, porcine hemagglutinating encephalomyelitis virus, porcine deltacoronavirus), and the zoonotic coronavirus infections (SARS and MERS coronaviruses).
Coronaviruses of Birds
INFECTIOUS BRONCHITIS VIRUS
Infectious bronchitis was the term coined in 1931 to describe the principal clinical-pathological feature of a transmissible respiratory disease of chickens in the United States first reported in North Dakota. Infectious bronchitis virus was identified retrospectively as the cause of a disease that had been misidentified as high-pathogenicity avian influenza in New England and the upper Midwest during 1924–1925. The disease has now been identified worldwide and is one of the most important viral diseases of chickens. The virus is the prototype of the genus Gammacoronavirus; there are many genotypes and serotypes as a consequence of mutations in its large genome.
Clinical Features and Epidemiology
The clinical presentation of infectious bronchitis depends on the age, genetic background, and immune status of the bird at the time of infection, route of exposure, nutritional factors (especially levels of calcium in the diet), virulence of the virus strain, and the presence of stressors such as cold temperatures, poor ventilation or secondary bacterial pathogens. Outbreaks may be explosive, with the virus spreading rapidly to involve the entire flock within a few days. The incubation period is typically brief: 18–48 hours. In chicks 1–4 weeks of age, virulent virus strains produce severe respiratory disease, with gasping, coughing, tracheal rales, sneezing, nasal exudate, wet eyes, respiratory distress, and, occasionally, swollen sinuses. Mortality in young chicks is usually 25–30%, but in some outbreaks can be as high as 75%. Less virulent strains cause fewer and milder respiratory signs, and lower morbidity and mortality rates. Infection of young female chicks may result in permanent hypoplasia of the oviduct that is evident later in life as reduced egg production and inferior quality eggs.
When the disease is uncomplicated by opportunistic bacterial superinfection, respiratory signs last for 5–7 days and disappear from the flock in 10–14 days. High mortality can occur in broilers as a result of secondary infection with Escherichia coli or pathogenic mycoplasmas. Egg-laying chickens usually present with reproductive tract involvement that is manifest as a decline or cessation in egg production or, less consistently, respiratory disease. When laying resumes, many eggs are abnormal, including lack of calcified shell, thin shells, and shells with stipples, distortions, dimples, depressions, or ridging; eggs that should be colored are often pale or white, and egg albumen may be watery. In acutely infected birds, the kidneys can be pale and swollen, with urates distending the ureters, and in the chronic phase there can be atrophy of kidney lobules, with large calculi within the ureters (urolithiasis).
Infectious bronchitis virus spreads between birds by aerosol and by ingestion of food contaminated with feces. In the environment, the virus can survive on fomites for several days and possibly for weeks, especially at low environmental temperatures. Outbreaks of infectious bronchitis have declined in recent years as a result of the extensive use of vaccines; however, the disease may occur even in vaccinated flocks when immunity is waning, or upon exposure to variant virus serotypes, with the first variant strains emerging in the 1940s and new variants continue to emerge today. To minimize this risk, most poultry producers obtain 1-day-old chicks from maternal antibody-positive breeders and then spray-vaccinate them with live-attenuated vaccine in the hatchery, with additional boosts by live-attenuated and/or inactivated vaccines. The current trend in “free-range” and backyard poultry production is likely to lead to a resurgence of infectious bronchitis.
Pathogenesis and Pathology
The virus replicates to high titer first in the respiratory tract (ciliated epithelial cells); this is followed by viremia (within 1–2 days of infection), which distributes the virus to many organs. The virus can cause extensive damage to the ovaries, oviduct, and the kidneys, but this is dependent on the properties of individual virus strains. The intestinal tract is another site of primary infection, but damage usually is minimal.
Infectivity declines rapidly, and isolation of virus beyond 7 days after infection is uncommon (except from chicks). Rarely, virus has been reported to persist for up to 14 weeks in cecal tonsils, and has been recovered from the feces for up to 20 weeks after infection. Kidney and intestine are the likely sites of virus persistence.
The most frequent gross pathologic finding is mucosal thickening within the upper and lower respiratory tract, with serous or catarrhal exudate in the nasal passages, trachea, bronchi, and airsacs. In very young chicks, the main bronchi may be blocked with caseous yellow casts. Pneumonia and conjunctivitis occur in some cases. In laying birds, ova can be congested and sometimes ruptured, with free yolk in the abdominal cavity. Desquamation of respiratory epithelium, edema, epithelial hyperplasia, mononuclear cell infiltration of the submucosa, and regeneration occur in various combinations. Repair processes begin after 6–10 days, and are complete in 14–21 days. Some virus strains affect the kidney, causing interstitial nephritis.
Diagnosis
Tracheal swabs or fresh samples of trachea are most useful for virus detection or isolation. Direct immunofluorescence staining of tracheal tissue smears is useful in the diagnosis of early cases before secondary bacterial infection has occurred. For virus isolation, embryonated chicken eggs are inoculated via the allantoic sac route. Infectious bronchitis virus does not typically infect cells in culture, although primary chick kidney cells can propagate the virus. Changes suggestive of the presence of a coronavirus include congestion of the main blood vessels in the chorioallantoic membrane and embryo stunting, curling, clubbing of down, or urate deposits in the mesonephros. Identification of virus in the chorioallantoic membrane is usually done by immunofluorescence or immunohistochemical staining, or in allantoic fluid by serological methods, nucleic acid analysis, or electron microscopy. Isolates are usually typed and subtyped by serologic methods and nucleic acid analyses such as genotype-specific RT-PCR assays.
Immunity, Prevention, and Control
Infection induces IgM, IgG, and IgA antibodies. In immune laying hens, the ovum begins to acquire IgG antibody (some of it virus specific) from the blood about 5 days before the egg is laid. As it becomes surrounded with albumen during passage down the oviduct, the ovum acquires both IgM and IgA antibodies, which are transferred into the amniotic fluid about halfway through development. During the last third of embryonation, IgG enters the circulation from the yolk; antibody can inhibit virus replication at this time. The chick hatches with a circulating IgG level similar to that of the hen. IgG antibody is metabolized with a half-life of approximately 3 days and may persist for 3–4 weeks. The virus may survive until passive immunity declines to a level at which it can replicate again, at which time the chicken mounts an active immune response. However, the correlates of active immunity to infectious bronchitis virus are less certain. Neutralizing antibodies can prevent virus dissemination from the respiratory tract and block secondary infection of the reproductive tract and kidneys. The adaptive transfer of CD8 T lymphocytes protects chicks against infectious bronchitis virus challenge, suggesting a role for cellular immunity as well in protection.
Live-attenuated virus vaccines are widely used to protect meat chickens. These vaccine viruses are derived by serial passage in embryonated chicken eggs. They are administered in drinking water, by coarse spray, or by deposition on the conjunctiva (eye drops). The first vaccination is typically given in the hatchery when birds are 1 day old, and booster vaccination is given at 10–18 days. Passively acquired maternal immunity prevents respiratory infection and disease for the first 7 days. For layers or breeders, live-attenuated vaccines are used for priming, followed by killed oil-adjuvanted booster vaccines, often given repeatedly during the laying cycle. Vaccination breaks occur because of the variable presence of new antigenic variants and existence of several serotypes. Such variants will continue to emerge and spread, posing continuing problems for poultry producers.
Control of infectious bronchitis is difficult because of the presence of persistently infected chickens in some flocks and the continuing emergence of antigenically variant viruses. The domestic chicken is the primary and most important host, but infections and disease have been described in pheasants infected with a closely related coronavirus. Sporadic or individual cases of avian infectious bronchitis virus infection also have been described in peafowl, teal, partridge, and guinea fowl. Avian coronaviruses related to infectious bronchitis virus have also been identified in many wild bird species, but these are typically found in the gastrointestinal tract.
TURKEY CORONAVIRUS
Coronaviruses were first recognized in turkeys in the United States in 1951 and were associated with various enteric disease syndromes, variously termed “blue comb disease,” “mud fever,” “transmissible enteritis,” and “coronaviral enteritis.” The disease is present throughout the world, essentially wherever turkeys are raised. The virus can infect turkeys of all ages, but the most severe enteric disease is evident within the first few weeks of life. The onset is characterized by loss of appetite, watery diarrhea, dehydration, hypothermia, weight loss, and depression. Younger poults may die. The duodenum and jejunum are pale and flaccid, and the ceca filled with frothy, watery contents. The feces may be green to brown, watery, and may contain mucus and urates. The cloacal bursa is small (atrophic). Some turkeys may shed virus in their feces for up to 7 weeks, with virus transmission by the fecal–oral route. Turkey coronavirus infections also result in reduced egg production in breeder hens, and eggs may lack normal pigment and have a chalky shell surface. Interaction between turkey coronavirus and other agents (E. coli, astrovirus, etc.) accentuate the disease.
Only one serotype of turkey coronavirus is recognized. Turkey coronavirus is classified, along with other avian coronaviruses, as a gammacoronavirus. Although there is high sequence identity (85–90%) in the three major viral proteins (polymerase, M, and N) of turkey coronavirus and avian infectious bronchitis virus, their S proteins are quite different, and turkey coronavirus likely represents a recombinant coronavirus containing a spike gene of unknown origin. Whether the origin of turkey coronavirus reflects altered enteric tropism or adaptation of an infectious bronchitis-like virus to the turkey, or whether infectious bronchitis virus is in itself a variant of an ancestral enteric avian coronavirus, is also unclear. Recently, bovine coronavirus was shown experimentally to infect turkey poults, but natural cases have not been described.
Turkey coronavirus can also be isolated in embryonated eggs of turkeys and chickens using the amniotic route of inoculation. No licensed vaccines for turkey coronavirus are available. Treatment involves supportive care, and is not specific.
Other CORONAVIRUSES OF BIRDS AND BATS
Warm-blooded flying vertebrates likely serve as the definitive hosts that harbor the coronavirus gene pool, with alpha- and betacoronaviruses having their origin in bats, and gamma- and delta-coronaviruses having their origin in birds. A wide variety of coronaviruses have been identified in geese, guinea fowl, swans, gulls, shorebirds, vulture, sparrow-hawk, hawk, woodpecker, fruit crow, great kiskadee, ruddy turnstone, pigeons, ducks, parrots, and other species of birds. Similarly, genetically divergent species of coronaviruses have been identified in a wide variety of species of bats, implying that they, like birds, may be the source of future epidemics of human and/or animal disease.
CORONAVIRUSES of Cats, Dogs, and Ferrets
FELINE ENTERIC CORONAVIRUS AND FELINE INFECTIOUS PERITONITIS VIRUS
Feline infectious peritonitis was first described in the 1960s as a systemic and often fatal disease of cats. The pathogenesis of feline infectious peritonitis is complex and not fully characterized, despite intensive study. Feline enteric coronavirus infection is central to the pathogenesis of this disease, as the sporadic occurrence of feline infectious peritonitis is proposed to be the result of mutations of the enteric coronavirus during natural infection of cats, resulting in the emergence of a virus with an acquired tropism for macrophages. Although all feline enteric coronaviruses are classified as alphacoronaviruses (Table 24.1), two distinct serotypes of the virus have been identified, both being able to cause feline infectious peritonitis. The majority of circulating feline coronaviruses are designated as serotype I. The serotype II feline enteric coronaviruses appear to be relatively rare, and represent recombinants that include portions of the genome of canine coronavirus, presumably arising from coinfection situations of feline and canine coronaviruses. Serotype II feline infectious peritonitis viruses grow well in cell culture and utilize amino-peptidase-N (APN) as a receptor. In contrast, serotype I viruses are very difficult to culture, and appear to use a distinct and currently unidentified receptor. However, both virus types can cause the two clinical forms of feline infectious peritonitis, one that has a characteristic abdominal effusion (the “wet” form), and the other (the “dry” form) without abdominal effusion. Thus, the pathologic manifestations are not solely a virus strain-specific property, as individual virus strains can cause either form of the disease in individual cats.
Clinical Features and Epidemiology
Feline infectious peritonitis is a common progressive, debilitating and lethal disease of domestic and wild members of the family Felidae. Disease typically occurs in young or very old cats, or in the context of immune suppression. The initial clinical signs are vague, and affected cats present with anorexia, chronic fever, malaise, and weight loss. Ocular and/or neurological manifestations occur in some individuals. In the classical wet or effusive form of feline infectious peritonitis, these signs are accompanied by progressive abdominal distention from the accumulation of a highly viscous fluid in the peritoneal cavity and rapid disease progression, with death typically within weeks to months. The dry or noneffusive form of the disease, with little or no peritoneal exudate, is more slowly progressive. The wet and dry forms of feline infectious peritonitis are different manifestations of the same infection, and both forms of the disease are characterized by foci of pyogranulomatous inflammation in several organs.
The following is a proposed scenario of fatal feline infectious peritonitis. A kitten suckling a seropositive queen is protected by colostral antibody against enteric coronavirus infection during the first few weeks of life. As maternal antibody wanes, the kitten becomes infected during an episode of maternal shedding of feline enteric coronavirus. The kitten now develops an active immune response, but in most cases not a sterilizing response, and a persistent viral infection of the gut with chronic fecal shedding is established. Virus and antibodies coexist in the kitten, but the infection is modulated by an efficient cellular immune response that keeps virus replication in infected macrophages and monocytes in check. The animal may remain healthy, but becomes susceptible to development of feline infectious peritonitis should it become stressed or immunosuppressed. Viral mutants then emerge, with rapid selection and proliferation of macrophage-tropic variants that cause the development of feline infectious peritonitis.
Pathogenesis and Pathology
The key initiating pathogenic event in feline infectious peritonitis is the productive infection of monocytes and macrophages by genetic variants (mutants) of the original enteric coronavirus. Experimentally, the virulence of strains of feline enteric coronavirus has been correlated with their capability of productive infection of cultured peritoneal macrophages, with avirulent strains infecting fewer macrophages and producing lower virus titers than virulent strains. Avirulent strains are also less able to sustain virus replication and spread between macrophages. Mutations within the spike (S) and, potentially, other proteins alter the tropism of the ubiquitous avirulent feline enteric coronavirus to macrophages, which then allows the virus to spread and ultimately to cause feline infectious peritonitis. The most consistently occurring mutations appear to be within the cleavage-activation and fusion domains of spike, and within the 3C accessory gene. Affected cats typically produce a strong antibody response that is ineffective in eliminating the virus, and cellular immune responses are unable to prevent virus replication in macrophages.
The lesions in feline infectious peritonitis are characteristically centered on small blood vessels, and vascular injury and leakage are central to the pathogenesis of the wet form of the disease. However, there is uncertainty regarding the pathogenetic mechanisms involved, as there is increasing evidence that vascular injury is not simply the result of immune complex deposition in the walls of the affected vessels, as was once proposed. The central role of viral infection of macrophages, however, is clear, and perivascular clusters of virus-infected macrophages are characteristically present in the tissues of cats with both the wet and dry forms of feline infectious peritonitis. Despite the inability of macrophages to prevent virus from replicating in them, infection of macrophages probably leads to their activation, with production of inflammatory mediators including cytokines and arachidonic acid derivatives (leukotrienes and prostaglandins). These mediators probably contribute substantially to the disease process, as these host-response molecules induce changes in vascular permeability and provide chemotactic stimuli for neutrophils and monocytes that further contribute to the inflammatory response. Both intravascular and recently emigrated monocytes and macrophages probably serve as new virus targets, thereby amplifying the infection further. The end result is enhanced local virus production, increased tissue damage, and a strong but ineffective host immune response.
Humoral immunity is not protective, and may actually enhance disease progression. Antibody-dependent enhancement of infection of macrophages is apparently mediated by neutralizing antibodies to the S protein, making vaccine development problematic. Cats that are seropositive to feline enteric coronavirus, either from natural infection or via purified IgG antibodies transfused into uninfected animals, develop an accelerated, fulminant disease when challenged experimentally with virulent feline coronavirus (so-called feline infectious peritonitis virus). Clinical signs and lesions develop earlier, and the mean survival time is reduced as compared with seronegative cats.
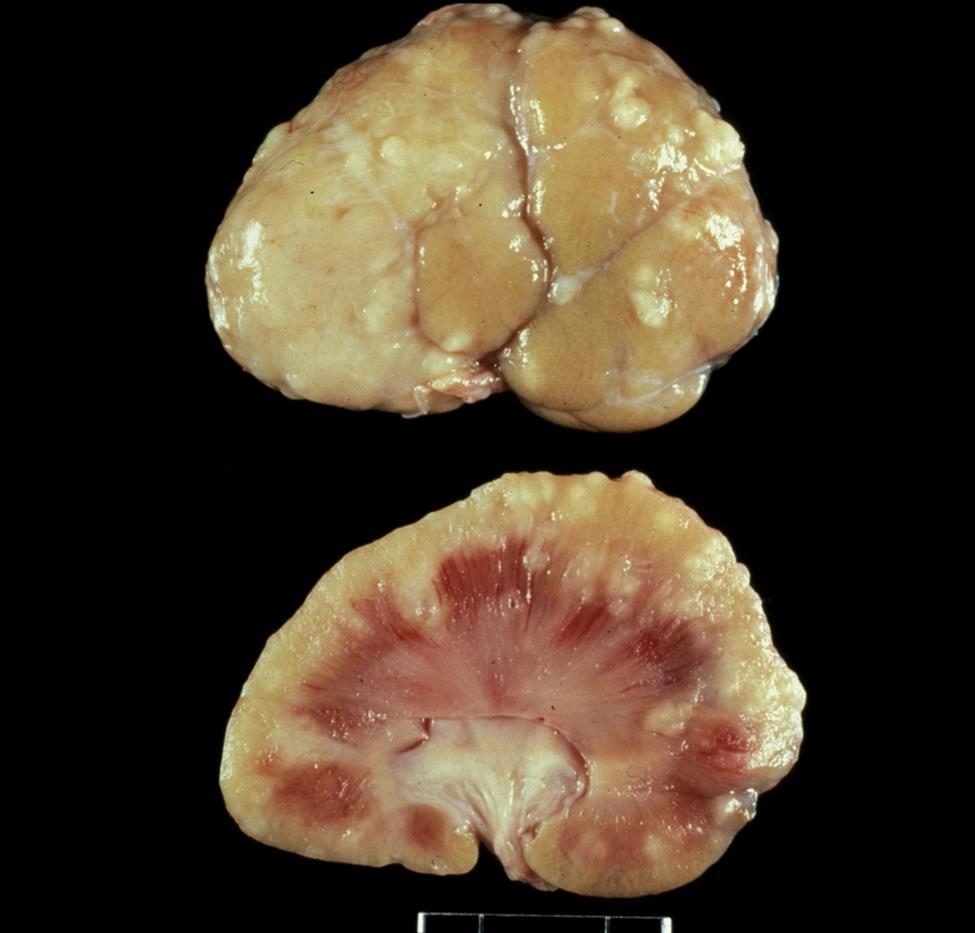
The gross lesions of feline infectious peritonitis reflect one of the two forms of the disease. The wet form is characterized by the presence of variable quantities of thick, viscous, clear yellow peritoneal exudate, and the presence of extensive fibrinous plaque with numerous discrete gray-white nodules (from <1 to >10 mm in diameter) in the omentum and on the serosal surface of the liver, spleen, intestines, and kidneys (Fig. 24.6). Microscopically, these nodules are composed of aggregates of macrophages and other inflammatory cells (granulomas or pyogranulomas) that characteristically are centered on blood vessels, sometimes with necrosis of the wall of involved vessels. These lesions can occur in many tissues, but omentum and peritoneal serosa, liver, kidney, lung and pleura, pericardium, meninges, brain, and uvea are common sites. The lesions and pathogenesis of the dry form of feline infectious peritonitis are similar, but without the fibrinous polyserositis that characterizes the wet form, and discrete pyogranulomas form nodular masses within the parenchyma of affected organs. It is unknown what determines the form of feline infectious peritonitis that develops in an individual cat; neither is the relationship between the two forms well understood, as individual virus strains can cause either form in different animals and both forms may be present in a single cat.
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree