Caliciviridae and Astroviridae
Abstract
This chapter describes the properties of caliciviruses and astroviruses, and features of the diseases they cause in animals.
Keywords
Calicivirus; astrovirus; Caliciviridae; Astroviridae; Norovirus; Lagovirus
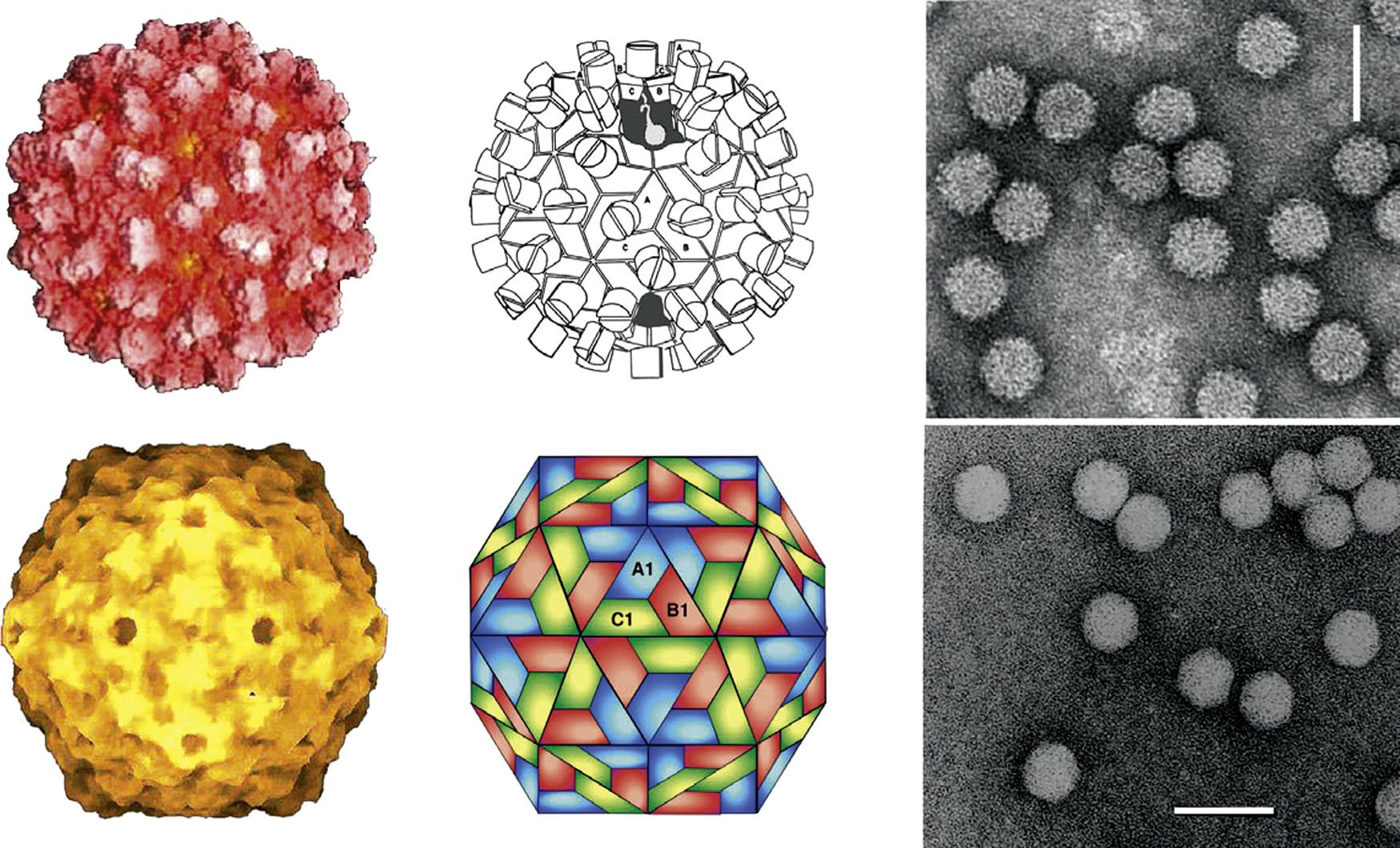
Members of the Caliciviridae and Astroviridae are biologically and taxonomically distinct, however they share some general features that make it convenient to consider them together. Caliciviruses and astroviruses are small, nonenveloped viruses with icosahedral capsids that enclose a single-stranded, positive-sense RNA genome—the capsid diameters and genome sizes of these viruses are similar, as is their overall capsid morphology. However, the surface appearance of their capsomers is different and can be distinguished by electron microscopy. Astrovirus virions may have a star-shaped (five- or six-points; hence the prefix astro-) surface appearance whereas calicivirus virions have cup-shaped surface depressions (calici is derived from the Latin calix for cup) or are fuzzy in appearance. The organization of the RNA genomes of caliciviruses and astroviruses is also similar and, like picornaviruses, both have a small viral protein, designated VPg, which is covalently bound at the 5′ end. The genomic RNA encodes two or three open reading frames.
The classification of member viruses of the families and Caliciviridae and Astroviridae, and their associated diseases, are discussed in the individual sections below. These viruses typically cause acute diseases with a short clinical course, although prolonged infections of individual animals occur with some of these viruses.
Members of the Family Caliciviridae
Members of the Caliciviridae infect a broad variety of animal species. Pathogenic caliciviruses typically cause enteric, oral cavity and upper respiratory, or systemic disease in their respective hosts. Although members of the family Caliciviridae have frequently proven difficult to isolate in cell culture, metagenomic approaches now allow new caliciviruses to be rapidly identified without the requirement for virus isolation. Calicivirus infections occur in humans, swine, birds, marine mammals, lagomorphs, rodents, monkeys, cattle, sheep, mink, cats, dogs, skunks, reptiles, fish, amphibians, and even insects, although the pathogenic significance of many of these viruses is currently uncertain.
Vesicular exanthema of swine is a disease of historical significance that was first recognized in southern California in 1932 and eradicated from domestic swine in the United States by 1956. Closely related caliciviruses have since been isolated from numerous species of marine mammals that occur throughout the northern Pacific Ocean, and are likely to have been the original source of swine infections. The major importance of vesicular exanthema of swine is that it mimics foot-and-mouth disease in swine. Feline calicivirus is a common cause of oral cavity and upper respiratory tract infection and disease in cats, and highly virulent systemic forms of the infection are also recognized. Rabbit hemorrhagic disease first emerged in China in 1984 as an apparently new, often fatal disease in domestic rabbits. This virulent virus likely arose from avirulent virus strains that circulated amongst European rabbit populations; and virulent rabbit hemorrhagic disease virus has since spread extensively and now is enzootic in rabbit populations in much of the world. A related calicivirus causes European brown hare syndrome. Murine caliciviruses (noroviruses) were discovered among laboratory mouse colonies. Infections are clinically silent, except in specific types of genetically altered (engineered) mice. The pathological significance of the noroviruses identified recently in many other species, including their zoonotic potential (if any), is currently uncertain.
Properties of CALICIVIRUSES
Classification
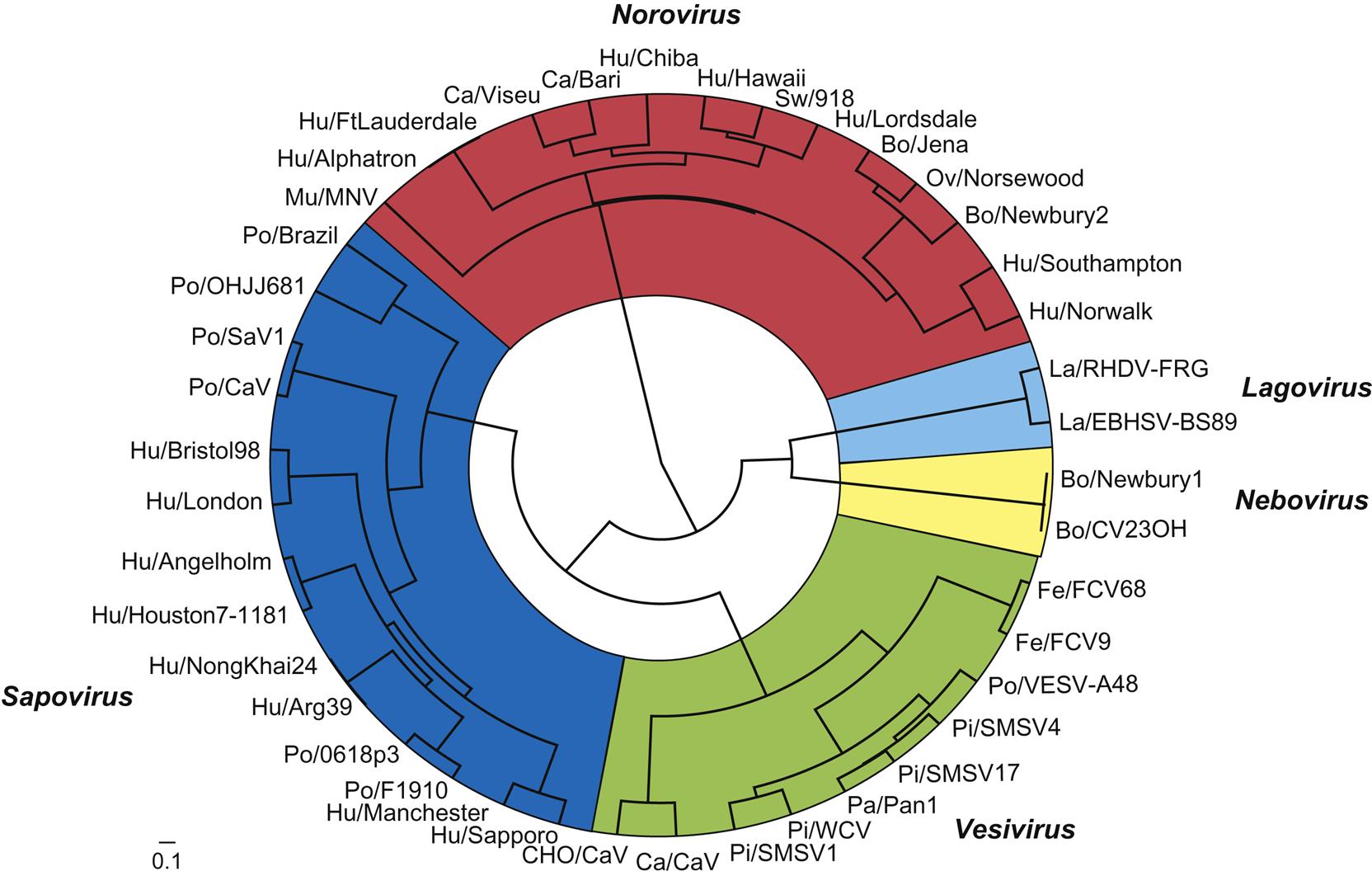
The family Caliciviridae currently comprises five genera (Vesivirus, Lagovirus, Norovirus, Sapovirus, and Nebovirus), with an additional four genera (Recovirus, Valovirus, Nacovirus, and Bavovirus) proposed. All genera include viruses that infect animals. The genus Vesivirus contains feline calicivirus and vesicular exanthema of swine virus, and close relatives that include the numerous San Miguel sea lion viruses, cetacean calicivirus, primate calicivirus, skunk calicivirus, bovine calicivirus, reptile calicivirus, and, tentatively, mink calicivirus. Several of these viruses share the ability to cause mucosal or cutaneous vesicles (“blisters”) in their respective hosts, thus their designation as vesiviruses. The genus Lagovirus (from Lagomorph) contains rabbit hemorrhagic disease and European brown hare syndrome viruses. The genus Norovirus contains six different genogroups with multiple subgroups within each. Noroviruses are associated with outbreaks of acute gastroenteritis in humans, and noroviruses that infect cattle, sheep, pigs, dogs, cats, and mice have been described. The genus Sapovirus also contains viruses that have been linked to outbreaks of human gastroenteritis and it tentatively now contains a number of strains of porcine enteric calicivirus, some of which are antigenically similar to human isolates, and a mink enteric sapovirus. The genus Nebovirus thus far contains two virus species that infect cattle, the type species Newbury-1 virus and bovine enteric calicivirus NB (Nebraska). These viruses cause mild enteritis in calves. Of the four other proposed genera, Recovirus, would contain Tulane calicivirus, a virus isolated from Rhesus monkeys (Macaca mulatta) with gastroenteritis, Valovirus would contain St-Valérien-like viruses that were isolated from pigs, and Nacovirus and Bacovirus would contain novel caliciviruses recovered from chickens and turkeys. With the advent of metagenomic sequencing of viruses from many species, the group of unassigned caliciviruses is growing rapidly.
The caliciviruses are genetically heterogeneous (Fig. 27.1). Sequence analysis of the capsid gene has identified at least five distinct genetic groups in both the genus Sapovirus and the genus Norovirus. Genetic recombination between viruses in the different genera complicates typing, and potentially might facilitate interspecies transmission of recombinant viruses, including from animals to humans. Additional viruses unquestionably will be added to this group as additional species are investigated.
Virion Properties
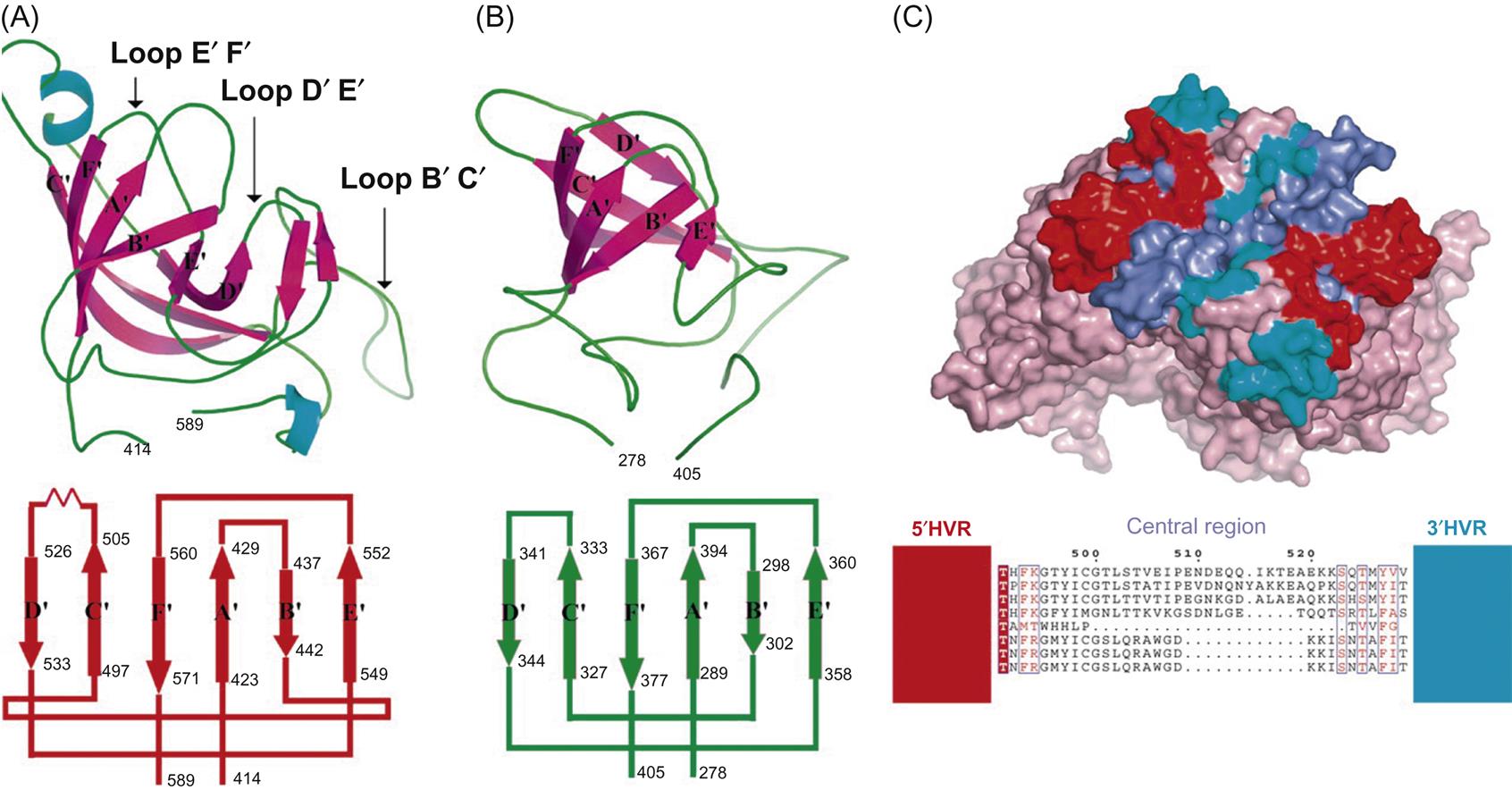
Calicivirus virions consist of a nonenveloped icosahedral capsid, 27–40 nm in diameter, which contains a positive-sense single-stranded RNA genome. The capsid is composed of 90 dimers of a single major capsid protein (VP1; 55–70 kDa) arranged as a T=3 icosahedral lattice (Fig. 27.2; Table 27.1). Each individual capsid protein is functionally divided into S (shell) and P (protrusion) domains. As the name implies, the P domain is the most externally exposed; it is further subdivided into P1 and P2 subdomains. The S domain is the most conserved and the P domain is most variable amongst virus strains. Analysis of the feline calicivirus capsid confirms that the hypervariable regions all lie within the P domain in loops that extend from the central conserved regions (Fig. 27.3). The neutralization epitopes and two linear B cell epitopes map to the hypervariable loops of the P2 subdomain, suggesting that the surface loops that are most accessible on the capsid are immunodominant and that changes in this region are responsible for antigenic variation. The receptor molecule for feline calicivirus, feline junctional adhesion molecule (fJAM-A) binds to the most accessible region of the protruding dimer. Feline calicivirus also contains a minor virion protein (8.5–23 kDa) that is essential for the assembly of infectious virions. The number of copies of this protein that are integrated into virions is low, but the exact number is not known.
Table 27.1
Virions are nonenveloped, 27–40 nm in diameter, with icosahedral symmetry
Some virions have a characteristic appearance, with 32 cup-shaped depressions on their surface
Virions are assembled from one capsid protein (Mr 60,000)
Genome is composed of a single molecule of linear positive-sense, single-stranded RNA, 7.4–8.3 kb in size
Genomic RNA is polyadenylated at its 3′ end and has a protein linked covalently to its 5′ end; genomic RNA is infectious
Cytoplasmic replication. Genomic RNA and several subgenomic mRNAs are produced during replication; mature proteins are produced both by processing of a polyprotein and by translation of subgenomic mRNAs
Currently five genera, with others proposed: Lagovirus, Nebovirus, Norovirus, Sapovirus, and Vesivirus
Because caliciviruses, like astroviruses, lack a lipid envelope, they are resistant to lipid solvents and standard detergent-based disinfectants. Caliciviruses are unstable at pH 6 or below, or pH 9 or above, and are inactivated by a solution of 10% sodium hypochlorite.
Virus Replication
Details of the replication cycles of the members of the family Caliciviridae have mostly been extrapolated from data for feline calicivirus and murine norovirus, which are both readily propagated in cell culture. Caliciviruses infect cells via receptor-mediated attachment and clathrin-dependent endocytosis, and replication then occurs on the surface of membranous vesicles in the cytoplasm of infected cells. Feline junctional adhesion molecule-A (fJAM-A) is a functional receptor for feline calicivirus, and the tissue distribution of the fJAM-A molecule explains the tropism of feline calicivirus infection in cats. Similarly, the distribution of various cell surface glycans that serve as attachment factors or receptors for murine norovirus likely determines the tissue tropism of this virus.
Members of the genus Vesivirus readily replicate in a variety of cultured cells and cause overt cytopathology as a consequence of apoptosis of infected cells. Other factors may be required for virus replication in cell culture; for example, replication of porcine enteric calicivirus (genus Sapovirus) in porcine kidney cells requires the presence of bile acids that increase levels of cyclic AMP in the treated cells, resulting in a downregulation of signal transducer and activator of transcription 1 (STAT1). This molecule (STAT1) is a key regulator of innate immunity and interferon response genes, and virus replication is enhanced in cells deficient in STAT1 and interferon receptor genes. Notably, clinical disease occurs in murine norovirus-infected, genetically engineered mice with STAT1 deficiency, but not in immunocompetent mice. The murine noroviruses are the first and only noroviruses to be successfully cultivated in vitro, and have therefore attracted much interest because of the importance of human norovirus infections. B cells and macrophages are likely targets for murine noroviruses in vivo. Although human norovirus infection causes enteric disease, recent findings suggest that B cells may also be primary cellular target.
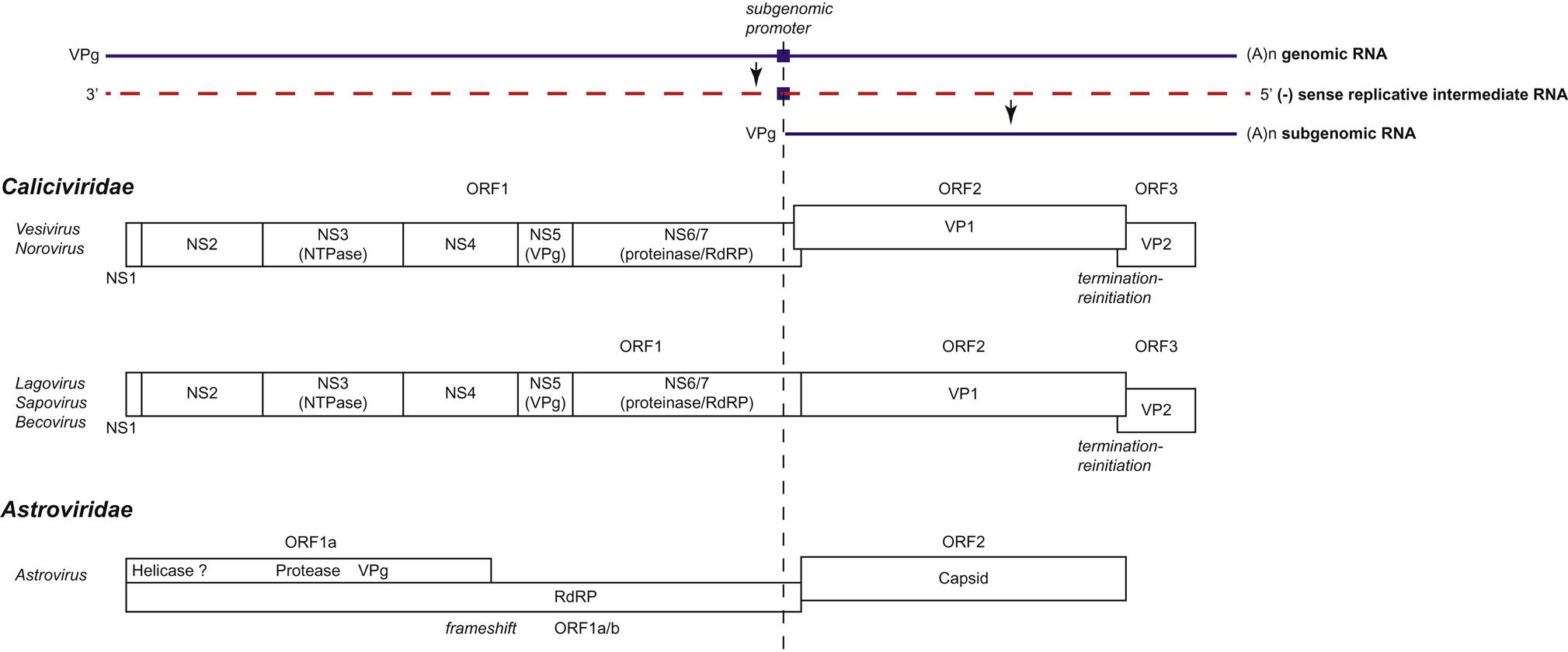
Individual caliciviruses utilize different strategies to generate the proteins essential for their replication (Fig. 27.4). The genome of caliciviruses includes either two or three open reading frames, with the 5′ end of the genome encoding the nonstructural proteins (helicase, VPg, protease, RNA polymerase) and the 3′ portion encoding the major and minor capsid proteins. Virus replication begins following the delivery of the genome into the cytoplasm of a host cell, and the viral genomic RNA is recognized by host ribosomes as an mRNA. The open reading frame (ORF1) closest to the 5′ end is the first to be translated, and individual nonstructural viral proteins, including the RNA-dependent RNA polymerase, are produced by autocatalytic posttranslational cleavage of the polyprotein encoded by ORF1 by a virus-encoded protease. In addition to full-length genomic RNA, a subgenomic positive-sense RNA coterminal with the 3′ end of the genome is present in infected cells; the VPg protein is covalently linked to the 5′ end of both the genomic and subgenomic RNA molecules, and this protein binds initiation factors for the translation of the viral RNAs. Translation of ORF3 from the subgenomic RNA of caliciviruses occurs as a result of ribosomal termination-reinitiation. The subgenomic RNA may function as a mechanism for the control of the level of translation of the structural proteins. Replication is associated with, and causes reorganization of, intracellular membranes derived from the endoplasmic reticulum. Virions accumulate in the cytoplasm, either scattered as paracrystalline arrays or as characteristic linear arrays associated with the cytoskeleton. Caliciviruses have no defined method of egress and are released by cell lysis, presumably after apoptosis of infected cells.